OEMolAdaptor2
Attention
This API is currently available in C++ and Python.
class OEMolAdaptor2 : public OEMolPotential:: OEMolAdaptor, public OEMolPotential:: OEMolFunc2
The OEMolAdaptor2 class is a pure virtual base class which defines the interface for classes which adapt or transform molecular coordinates, gradients, and hessians of a given OEMolFunc2 during an optimization. Adaptors are useful in cases where energy, gradients and hessians evaluation for a molecule or conformation are performed in Cartesian coordinates, but the optimization is done in different coordinate system.
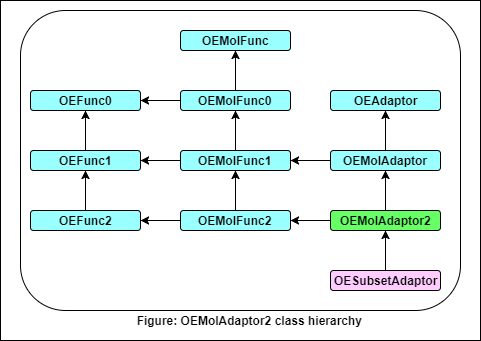
- The following methods are publicly inherited from OEAdaptor:
- The following methods are publicly inherited from OEFunc0:
- The following methods are publicly inherited from OEFunc1:
- The following methods are publicly inherited from OEFunc2:
- The following methods are publicly inherited from OEMolFunc:
- The following methods are publicly inherited from OEMolAdaptor:
- The OEMolAdaptor2 class defines the following public methods:
- The following classes derive from this class:
SetMolFunc2
bool SetMolFunc2(OEMolFunc2 &molfunc, bool own)=0
Defines the interface for resetting the molecule function, OEMolFunc2, within a
OEMolAdaptor2 derived instance. Any molecule functions which are
“owned” at the time of calling this method will be deleted. If the value of a second argument
passed to the method is true, the adaptor derived from OEMolAdaptor2
will take ownership of the memory of the molecule function, and will call the molecule function’s
delete operator when either the destructor or the current method is called.