OEQuatAdaptor
Attention
This API is currently available in C++ and Python.
class OEQuatAdaptor : public OEMolAdaptor
The OEQuatAdaptor class provides the ability to rotate and translate a molecule as single rigid unit or solid body in the presence of an external field. Rotations are represented with quaternion variables.
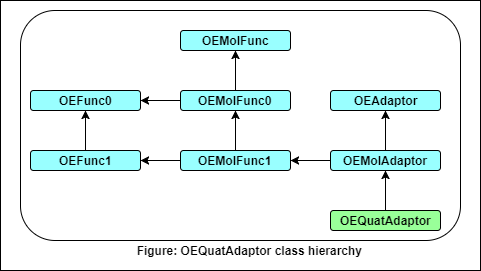
- The following methods are publicly inherited from OEAdaptor:
- The following methods are publicly inherited from OEFunc0:
- The following methods are publicly inherited from OEFunc1:
- The following methods are publicly inherited from OEMolFunc:
- The following methods are publicly inherited from OEMolAdaptor:
- The OEQuatAdaptor class defines the following public methods:
Constructors
OEQuatAdaptor(OEMolFunc1 &, bool own=false, bool excludeInteract=true)
Default and copy constructors.
Constructs an OEQuatAdaptor instance. The molecule function used
for function evaluation must be provided as the first argument to the constructor. The
second argument specifies whether the OEQuatAdaptor object takes
ownership of the memory of the molecule function instance. By default that does not happen,
so the OEQuatAdaptor destructor does not delete the molecule function
instance. If ownership of the molecule function is transferred to the
OEQuatAdaptor instance, the molecule function’s delete operator will
be called in the OEQuatAdaptor destructor. The third argument
controls the inclusion of invariant molecule function interactions. By default, interactions
which remain constant during the solid-body optimization (all intramolecular interactions)
are excluded from calculation. If the third calling argument is false then all
interactions will be computed during every function evaluation.
AdaptGrad
bool AdaptGrad(double *qg, const double *g, const double *quat,
const double *coords) const
Transforms a set of the input Cartesian gradients (second argument) into rotation and translation gradients which are copied to the first argument array. Third and fourth arguments are the arrays of quaternion plus translation vector and Cartesian coordinates respectively, used for the transformation.
IsRedundant
bool IsRedundant(const double* x, const double* y, const double tolerance=0.1) const
Compares two sets of quaternions-translational vectors coordinates. If points x and
y represent the same point in Cartesian space within given tolerance, method returns
true.