Ring Perception
Cycle Membership
The simplest form of ring processing in OEChem TK is testing
whether an atom or bond is in a ring or not.
The OEChem TK function
OEFindRingAtomsAndBonds is
used to determine which atoms and bonds are members of one or more
rings and which are acyclic. This function uses an efficient
\(O(n)\) algorithm. Once
OEFindRingAtomsAndBonds has
been called, an atom or bond can be tested for being in a ring by
calling the OEAtomBase::IsInRing or the
OEBondBase::IsInRing methods respectively.
The function
OEFindRingAtomsAndBonds is
called automatically by the high-level file read function
OEReadMolecule and
OESmilesToMol.
However, whenever you modify a molecule by adding or deleting bonds, you’ll need to
explicitly call OEFindRingAtomsAndBonds.
The following two ‘equivalent’ code snippets demonstrate
how to loop over chain
atoms using the OEAtomIsInRing
functor and the IsInRing
method of the OEAtomBase class.
for (OEIter<const OEAtomBase> atom = mol.GetAtoms(OENot<OEAtomBase>(OEAtomIsInRing())); atom; ++atom)
cout << atom->GetIdx() << ' ' << OEGetAtomicSymbol(atom->GetAtomicNum()) << endl;
for (OEIter<const OEAtomBase> atom = mol.GetAtoms(); atom; ++atom)
{
if (!atom->IsInRing())
cout << atom->GetIdx() << ' ' << OEGetAtomicSymbol(atom->GetAtomicNum()) << endl;
}
The chain/ring bonds of a molecule can similarly be accessed using the
OEBondIsInRing functor and the
IsInRing method of the
OEBondBase class.
For more information about functors see chapter Predicate Functors.
The user can also set the atom and bond ring flags manually using the
OEAtomBase::SetInRing and
OEBondBase::SetInRing methods.
Code Example
Ring Perception OpenEye Python Cookbook recipe
Membership in a Given Ring Size
It is also possible to use OEChem TK to determine whether an atom or a bond is in a ring
of a given size, using the OEAtomIsInRingSize
and OEBondIsInRingSize functions.
Both of these functions require that
OEFindRingAtomsAndBonds has previously been
called on the molecule.
Both of these functions take the query ring size as an argument, which should
be greater than or equal to three.
The definition of ring or cycle is not based upon the ‘’smallest set of smallest rings’’
(SSSR), and the functions return true if there is a bonded path of size unique atoms
where each atom is bonded to the next and the last is bonded to the first.
It is often the case that atoms may be in different sized cycles at the same time. For example, one way to identify the ring fusion atoms in indole (the fusion of a five-membered pyrrole ring and a six-membered benzene ring) is the following:
for (OEIter<OEAtomBase> atom = mol.GetAtoms(); atom; ++atom)
{
OEAtomBase* aptr = atom;
if (OEAtomIsInRingSize(aptr, 5u) && OEAtomIsInRingSize(aptr, 6u))
cout << aptr->GetIdx() << endl;
}
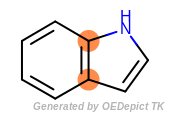
The fused atoms in indole belong to both a five- and a six-membered rings
OEChem TK also provides an additional pair of functions,
OEAtomIsInAromaticRingSize and
OEBondIsInAromaticRingSize, to
determine whether an atom or bond is in an aromatic ring or cycle of a given size.
These behave identically to OEAtomIsInRingSize
and OEBondIsInRingSize except that each ring bond
in the path/cycle must be aromatic.
In addition to OEFindRingAtomsAndBonds,
these functions also require the user to have called
OEAssignAromaticFlags.
Code Example
Ring Perception OpenEye Python Cookbook recipe
Smallest Ring Membership
In addition to determining whether an atom or a bond is in a ring or cycle of
a given size, it’s often useful to know the size of the smallest ring or cycle
that an atom or bond is in.
To do this OEChem TK provides the functions
OEAtomGetSmallestRingSize and
OEBondGetSmallestRingSize.
For acyclic atoms and bonds, these functions return the value zero.
For cyclic atoms and bonds, they return a value greater than or equal to
three.
for (OEIter<OEAtomBase> atom = mol.GetAtoms(); atom; ++atom)
{
OEAtomBase* aptr = atom;
const auto size = OEAtomGetSmallestRingSize(aptr);
if (size == 0u)
cout << aptr->GetIdx() << " acyclic" << endl;
else
cout << aptr->GetIdx() << " smallest ring size = " << size << endl;
}
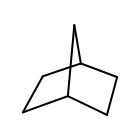
OEAtomGetSmallestRingSize returns five for each heavy atom of norbornane
Code Example
Ring Perception OpenEye Python Cookbook recipe
Connected Components Identification
To aid in splitting molecules into discrete connected components, for
example to separate a parent compound from its salt, or a ligand from
a protein, OEChem TK provides the function
OEDetermineComponents.
This function arbitrarily assigns an integer index, starting from one, to
each disconnected part in the OEMolBase.
The user provides a pointer to an array of at least
GetMaxAtomIdx unsigned ints,
which is populated by the function.
On return this provides a mapping from each atom’s index, obtained by
OEAtomBase::GetIdx, to its component index.
Unused atom indices are mapped to zero.
The function itself also returns the total number of components found, i.e.
the maximum part index stored in the array.
The following snippet provides a short example of how to use this function.
void MoleculeParts(const OEMolBase& mol)
{
vector<unsigned int> parts(mol.GetMaxAtomIdx());
const auto count = OEDetermineComponents(mol, &parts[0]);
printf("The molecule has %d components\n", count);
for (OEIter<const OEAtomBase> atom = mol.GetAtoms(); atom; ++atom)
printf("atom %d is in part %d\n", atom->GetIdx(), parts[atom->GetIdx()]);
}
Code Example
Splitting multicomponent molecules OEChem TK example
Ring Perception OpenEye Python Cookbook recipe
Ring Systems Identification
The OEChem TK function OEDetermineRingSystems
and OEDetermineAromaticRingSystems
behave very similarly to the OEDetermineComponents.
However, these functions return a mapping from atom indices to a ring or aromatic ring
system index, respectively (see example in Listing 1).
These functions require that OEFindRingAtomsAndBonds
has been called previously.
The function OEDetermineAromaticRingSystems also requires
the aromaticity perception by calling the OEAssignAromaticFlags function.
The OESmilesToMol used in the above example automatically calls
both the OEFindRingAtomsAndBonds and OEAssignAromaticFlags
functions.
When using the OEDetermineRingSystems function,
all acyclic atoms are mapped to the value zero.
When using the OEDetermineAromaticRingSystems
function, all aliphatic atoms are mapped to the value zero.
Listing 1: Aromatic ring system identification
#include <openeye.h>
#include <oeplatform.h>
#include <oesystem.h>
#include <oechem.h>
using namespace std;
using namespace OEPlatform;
using namespace OESystem;
using namespace OEChem;
int main()
{
OEGraphMol mol;
OESmilesToMol(mol, "C(O)(=O)c1cccc2c1[nH]c(C3CCCc4c3cccc4)c2");
vector<unsigned int> parts(mol.GetMaxAtomIdx());
const auto numaromsystems = OEDetermineAromaticRingSystems(mol, &parts[0]);
cout << "Aliphatic atoms: ";
for (OEIter<const OEAtomBase> atom = mol.GetAtoms(); atom; ++atom)
{
if (parts[atom->GetIdx()] == 0u)
cout << atom->GetIdx() << ' ';
}
cout << endl << "Number of aromatic ring systems = " << numaromsystems << endl;
for (auto ringidx = 1u; ringidx <= numaromsystems; ++ringidx)
{
cout << ringidx << " . aromatic ring system: ";
for (OEIter<const OEAtomBase> atom = mol.GetAtoms(); atom; ++atom)
{
if (parts[atom->GetIdx()] == ringidx)
cout << atom->GetIdx() << ' ';
}
cout << endl;
}
return 0;
}
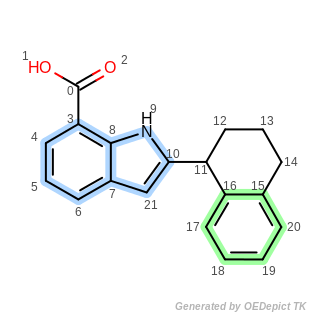
Example structure of aromatic ring system identification
The output of Listing 1 is the following:
Aliphatic atoms: 0 1 2 11 12 13 14
Number of aromatic ring systems = 2
1 . aromatic ring system: 3 4 5 6 7 8 9 10 21
2 . aromatic ring system: 15 16 17 18 19 20
Code Example
Ring Perception OpenEye Python Cookbook recipe
Smallest Set of Smallest Rings (SSSR) Considered Harmful
In graph-theoretical terms, a bond is considered cyclic if its removal from the structure would not lead to the structure being broken into separate components. Despite this simple definition, a large number of algorithms for ring detection exist. (See article [Downs-1989] for an extensive and comparative review.) The diversity and multitude of the ring perception methods derives from the fact that, while determining whether an atom is part of a ring is a very simple problem, the identification of “chemically meaningful” rings among the potentially large number of cyclic subgraphs of a molecular structure can be a surprisingly complex task.
The Smallest set of smallest rings (SSSR) [Plotkin-1971] is the most broadly used type of ring set in computational chemistry. However SSSR is not a unique subset of all possible cycles of a molecule (see Figure: Example of SSSR). Obviously SSSR membership can not be used as a graph theoretical invariant in symmetry perception. Indeed the choice of which rings are part of the SSSR and which are not is arbitrary, and often dependent upon the input order of the molecule. Because of the potential ambiguity of SSSR, many alternative ring set definitions to SSSR have been proposed over the years, including extended SSSR, the set of “synthetically important” rings, the set of elementary rings (SER), the essential set of essential rings (ESER), \(\kappa\)-rings, etc…
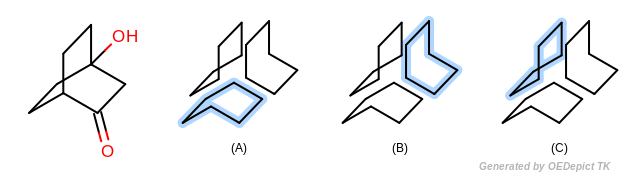
Example of SSSR SSSR is not an invariant subset of all possible rings; (a), (b) and (c) depict the three equally valid SSSR of the bridged structure (on the left)
We believe that it is a great service to our customers that we do not include any SSSR functionality in OEChem TK. This is a conscious (and consensus) decision. The forerunners of OEChem TK, Babel and OELib, both contained efficient algorithms for determining SSSR, and these remain freely available on the Internet today. Furthermore, many useful ring perception routines are available in OEChem TK, including:
the ability to detect whether an atom or bond is acyclic or part of a ring (For more details on
OEAtomBase::IsInRingandOEBondBase::IsInRing, see section Cycle Membership)the ability to detect whether an atom or a bond is in a ring of a particular size (For more details on
OEAtomIsInRingSizeandOEBondIsInRingSize, see section Membership in a Given Ring Size)the ability to detect whether an atom or a bond is in an aromatic ring of a particular size (For more details on
OEAtomIsInAromaticRingSizeandOEBondIsInAromaticRingSize, see section Membership in a Given Ring Size)the size of the smallest ring an atom or bond belongs to (For more details on
OEAtomGetSmallestRingSizeandOEBondGetSmallestRingSize, see section Smallest Ring Membership)the ability to determine whether atoms belong to the same ring system (
OEDetermineRingSystems) or the same aromatic ring system (OEDetermineAromaticRingSystems). See section Ring Systems Identification for more details.