OEToolkits 2017.Oct
Shape TK: Version 2.0.0
The 2017.Oct release introduces Shape TK version 2.0.0, which is completely redesigned with a new API that is more flexible and easily extended with new functionality. Shape TK version 2.0.0 is now thread-safe and is able to:
calculate color using grid- or proxy grid-based analytic methods
calculate shape overlap and color score between two fixed-in-space objects with a single calculation using the OEOverlapFunc class
combine force fields from the OEFF TK with shape and/or color for overlay optimization
obtain top hits from a database against a given query using the OEROCS class
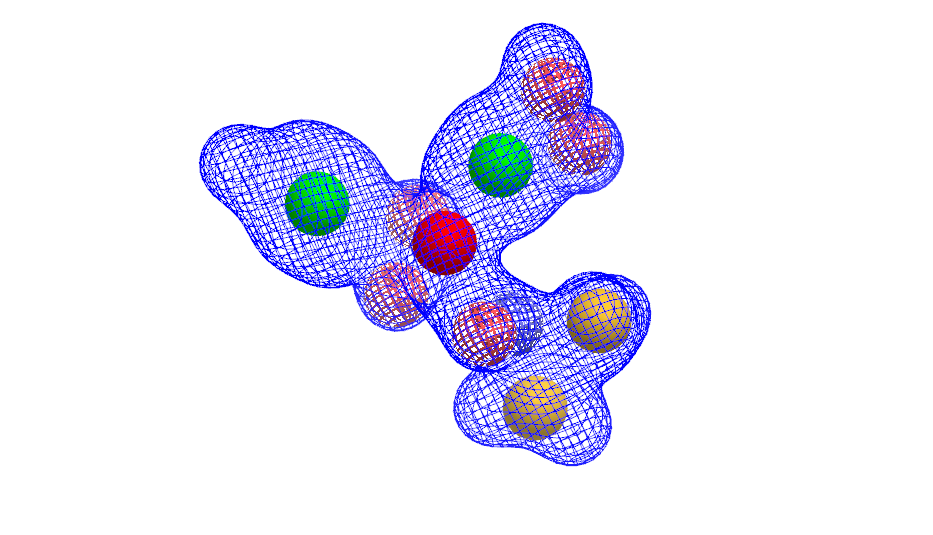
create a query combining any combination of a molecule, a grid, and multiple shape and/or color Gaussians using the OEShapeQuery class (see example below)
Preliminary testing of functionality from Shape TK version 2.0.0 indicates that overlay optimization is up to 10% faster than the 2017.Jun version.
|
Omega TK: New Torsion Library
A new torsion library has been added to Omega TK based on a probabilistic analysis of torsion angles in the Cambridge Structural Database (CSD) by Guba et al. [Guba-2016] This torsion library, which contains more than 500 entries, complements the existing torsion library, which is based mainly on analysis of structures found in the Protein Data Bank (PDB). Replacement of the existing library with the new library has four important effects on OMEGA’s performance:
nearly complete elimination of torsion values not well represented in the CSD
significantly improved run time
substantial reduction in the size of the conformer ensemble
small trade-off increase in the median RMSD for solid state structures (both from the PDB and the CSD)
OEDepict TK and Grapheme TK: Interactive SVG Images
The 2017.Oct release expands these toolkits’ ability to assemble sets of graphical objects in SVG images.
Images can present great deal of information very quickly, but those that try to convey too much information at once can look convoluted. The solution lies in generating interactive images that on the surface are simple but that can reveal extra information on request. This release allows users to mouse over simple images to view additional embedded information.
In the example below, the sequence-level depiction of a peptide is easily comprehensible due to its simplicity (see image below). The more detailed atomic-level representation of the amino acid components of the peptide can be revealed on mouse-over without cluttering the simplified view.
Hover the mouse over any peptide circle |
Hover the mouse over any residue circle or “Legend” label |
|
|
cyclic trypsin inhibitor (PDB: 1JBL) |
protein-ligand visualization (PDB: 3EQB) |
For more information, see the Generating Interactive SVG Images chapter of the OEDepict TK documentation.
General Notices
On macOS Python,
single-builddistributions are now installed by default when using the meta-installer viapip install openeye-toolkits.The Python verification tests now use
pytestrather thannosetestas the test discovery tool.pytestoptions can be passed to theopeneye.examples.openeye_testsmodule to control the tests and level of debugging information provided.This is the last release to support Java 1.6.
This is the last release to support Python 2.7.