Atom Association and Surface Splitting
Molecular and accessible surfaces are created with an atom associated to each
vertex. These atom indices can be obtained using the
OESurface::GetAtoms and
OESurface::GetAtomsElement methods, but the default
assignments are not very accurate. Using the
OESurfaceToMoleculeDistance function will ensure that each
vertex’s atom assignment is to the atom whose van der Waals surface is closest
to that vertex. While the associations provide an accurate mapping of
individual vertices to atoms, triangles on the surface may have vertices
belonging to different atoms. This can make make mapping of atom-based
properties onto the surface problematic.
Surface splitting, using the OESplitSurfaceByAtoms
function, will divide a surface such that the boundaries between atoms are
distinct, with no triangles being shared by multiple atoms. Surfaces are
split through a process of moving vertices and subdividing triangles until no
triangle edges cross the analytical boundaries between atoms. A comparison of
the atom associations of unsplit and split surfaces is shown in
Table: Atom Associations.
Atom Associations
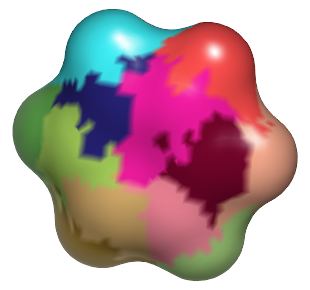
|
Molecular surface of benzene, colored by atom indices assigned during surface construction |
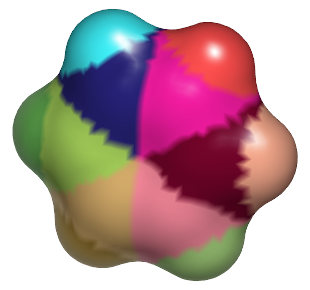

|
Same surface after atom indices are reassigned using OESurfaceToMoleculeDistance |
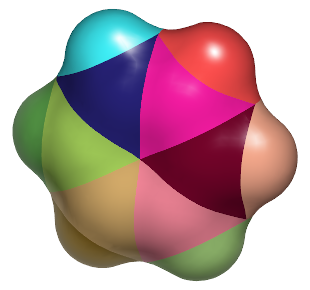

|
Same surface after splitting with OESplitSurfaceByAtoms |
Splitting a surface has some consequences that you should be aware of. A split surface will have significantly more vertices and triangles than the surface it was created from. In addition, it will contain numerous duplicate vertices, having the same coordinates and normals but with different atom associations. Finally, it will contain “zero-area” triangles, generated between duplicated vertices to preserve the connectivity of vertices within the surface.