Depicting CSV or SDF in HTML¶
Problem¶
You want to depict molecules along with their associated data read from a CSV file in an HTML file. See the generated HTML file in drugs.html and its screen-shot in Figure 1.
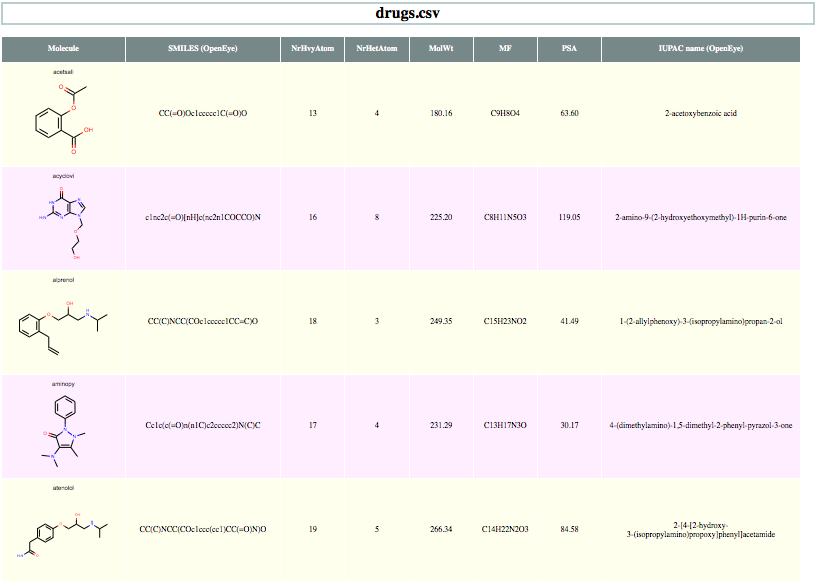
Figure 1. Example of depicting CSV in HTML (The screen-shot is reduced here for visualization convenience)
Ingredients¶
|
Solution¶
The CSV file format is a text file format containing comma-separated values. In OEChem TK this file format is implemented to enable data exchange with a wide variety of other software. Each line of a CSV file stores data for a molecule that is represented by a SMILES string.
See also
- CSV File Format section of the OEChem TK documentation about the layout of the CSV file format.
When reading a CSV file, the fields of the file are attached to each molecule as SD data. This data can be accessed by the OEGetSDDataIter function that returns an iterator over all the SD data (tag - value) pairs of a molecule. The CollectDataTags function iterates over a list of molecules and returns the unique tags of the data attached to the molecules.
1 2 3 4 5 6 7 8 9 | def CollectDataTags(mollist):
tags = []
for mol in mollist:
for dp in oechem.OEGetSDDataIter(mol):
if not dp.GetTag() in tags:
tags.append(dp.GetTag())
return tags
|
The WriteHTMLFile function takes a list of molecules read from a CSV file along with the data tags returned by the CollectDataTags function. It first writes the header of the html file, followed by iterating over the molecules and adding a new row into a table for each molecule by calling the WriteHTMLTableRow function. Finally, it finishes writing the html file by calling the WriteHTMLFooter function.
1 2 3 4 5 6 7 8 | def WriteHTMLFile(ofp, mollist, iname, tags, opts):
WriteHTMLHeader(ofp, iname, tags)
for mol in mollist:
WriteHTMLTableRow(ofp, mol, opts, tags)
WriteHTMLFooter(ofp)
|
The WriteHTMLHeader function sets the style of the html file (lines 5-13) and then adds the header of a table in which the molecules along with their data will be inserted (lines 21-28).
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 | def WriteHTMLHeader(ofp, filename, tags):
tablewidth = min(1800, (len(tags) + 1) * 200)
ofp.write("<style type='text/css'>\n")
ofp.write("h1 "
"{ text-align:center; border-width:thick; border-style:double;"
"border-color:#BCC; }\n")
ofp.write("table.csv "
"{ border-spacing:1; background: #FFF; width:%dpx; }\n" % tablewidth)
ofp.write("table.csv td, th "
"{ width:100px; text-align:center; padding:3px 3px 3px 3px; }\n")
ofp.write("table.csv th { height:50px; color:#FFF; background:#788; }\n")
ofp.write("table.csv tr:nth-child(even){ color:#000; background:#FFE; }\n")
ofp.write("table.csv tr:nth-child(odd) { color:#000; background:#FEF; }\n")
ofp.write("table.csv tr:hover { color:#000; background:#DDD; }\n")
ofp.write("</style>\n")
ofp.write("<html><h1>%s</h1>\n" % filename)
ofp.write("<body>\n")
ofp.write("<table class=csv>\n")
ofp.write("<tbody>\n")
ofp.write("<tr>\n")
ofp.write("<th> Molecule</th>")
# write data tags in table header
for tag in tags:
ofp.write("<th> %s </th>" % tag)
ofp.write("\n</tr>\n")
|
The WriteHTMLTableRow function inserts the image of the molecule (line 7) along with its corresponding data (lines 11-15) into the next row of the html table.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 | def WriteHTMLTableRow(ofp, mol, opts, tags):
ofp.write("<tr class=row>\n")
# add image
ofp.write("<td> %s \n </td>\n" % GetSVGImage(mol, opts))
# write data
for tag in tags:
value = "N/A"
if oechem.OEHasSDData(mol, tag):
value = oechem.OEGetSDData(mol, tag)
ofp.write("<td> %s </td>" % value)
ofp.write("</tr>\n")
|
The GetSVGImage generates and molecule display and returns its image as a string in bare svg image file format (with no header). This “image” string can be directly inserted into an html file.
1 2 3 4 5 6 | def GetSVGImage(mol, opts):
oedepict.OEPrepareDepiction(mol)
disp = oedepict.OE2DMolDisplay(mol, opts)
imagestr = oedepict.OERenderMoleculeToString("bsvg", disp, False)
return imagestr.decode("utf-8")
|
The WriteHTMLFooter function simply closes the table and the body of the html file.
Download code
csv2html.py and drugs.csv supporting data file
Usage
Running the above command will generate the drugs.html file.
prompt > python3 csv2pptx.py drugs.csv drugs.pptx
Discussion¶
Reading the columns of an CSV file into SD data fields, means that the OEChem TK provides a meta-data interchange between sdf files and CSV files. Consequently, the same Python script can be used to generate an html file reading an sdf file.
Usage
After downloading drugs.sdf supporting data file, the above command will generate the same drugs.html file (apart from the input filename on the top).
prompt > python3 csv2html.py drugs.sdf drugs.html
See also in OEChem TK manual¶
Theory
- SD Tagged Data Manipulation section
- CSV File Format section
API
- OEGetSDDataIter function
See also in OEDepict TK manual¶
Theory
- Molecule Depiction chapter
API
- OE2DMolDisplay class
- OE2DMolDisplayOptions class
- OEPrepareDepiction function
- OERenderMoleculeToString function
