Filtering Docking Hits by Interactions Tutorial¶
This tutorial and the floe described in it are intended to be used with output datasets containing docked poses from the OEDocking - Dock into an Active Site for Virtual Screening, the Gigadock Warp, or the Gigadock floes.
The example datasets used in the tutorial can be found in the /Organization Data/OpenEye Data/Filter_by_Interaction_hits folder.
For more on GigaDocking, see the tutorials _giga_dock_tutorial and _gigadock_warp_tutorial in the OpenEye-large-scale-floes package.
The “Filtering Docking Hits by Interaction Hints” floe requires two datasets as inputs: one for the receptor(s) used for docking, the other for the docked molecules. It then writes these same two datasets back out with an additional set of fields on each, which can be used to filter the docked poses by interactions with the target. Note that results generated using a single receptors or multiple receptors can be used in this floe.
The floe is solely for filtering of docked poses after docking, and not related to adding or enforcing constraints during the docking processes. The interactions between the target and ligand are perceived using _chapter_interactions from the OpenEye OEChem/OEBio toolkit. Additionally, an interaction fingerprint for each docked ligand in the receptor is calculated and a Tanimoto coefficient with the interactions from the bound ligand in the receptor as the reference (if available) is reported.
Floes used in the Tutorial¶
The floe used in this tutorial is documented here: Filter Docking Hits by Interactions Floe.
Running the Floe¶
To run the Filter Docking Hits by Interactions Floe, navigate to the Floe tab in Orion, then:
Search for Filter Docking Hits by Interactions in the search box or find it in the Classic Floes package.
Click the Floe to open the Job Form.
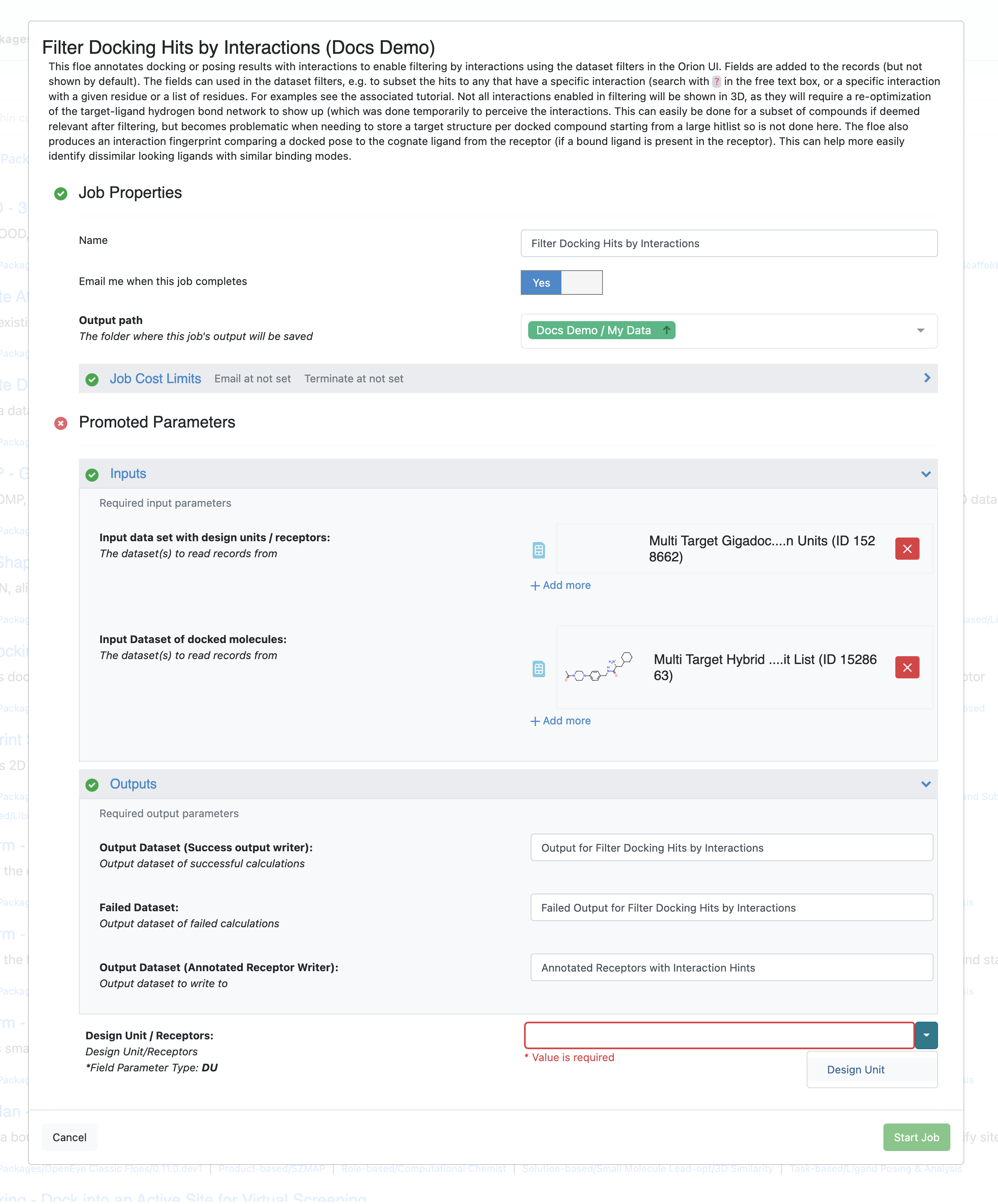
Running the Filter Docking Hits by Interactions Floe¶
The Floe takes two datasets: one containing the receptor(s) used to produce the docking hit list, the other containing the poses of the docked molecules (hit list). There is an additional required parameter for the floe where a user selects the name of the field containing the design units in the receptor dataset. The floe should automatically inspect the relevant datasets and suggest the correct field if the little arrow next to the field is clicked as shown in the image above.
Exploring the Interaction Fingerprints¶
In case a protein-ligand complex was used as receptor for docking, the output dataset contains a field called “InteractionFingerPrintSimilarity” which is a Tanimoto score (bounded between 0-1), comparing the similarity of a docked pose to the ligand that was bound in the docking receptor (if available) which in our case is the bound ligand from the crystal structures. The fingerprint is a bitvector constructed by multiplying the number of residues in the target binding site (aggregated if multiple receptors are used) with the number of perceived interactions. Then, for each ligand,a bit is turned on if the docked pose has an interaction with a given residue. This is similarly done for the ligand bound in the docking receptor, and a Tanimoto score of the similarity between the two fingerprints is calculated. Thus, if a docked pose has a high similarity, the compounds are not necessarily similar (not evaluated here, but can be done separately using molecular fingerprints or ROCS), but instead display similar binding modes, e.g. hydrogen bonds, salt-bridges, and contact interactions in the binding site.
Filtering the Docked Poses¶
First, we are going to mark both of the newly created datasets as activate:
Each of the fields added to the datasets for filtering is prefixed “Inter_”. These are hidden from the spreadsheet view by default, since they do not add value in the spreadsheet view and can be made visible if desired using the “Data Handling” menu.
The first filter we are going to apply only shows poses of molecules that have an intermolecular hydrogen bond to the receptor. This is done using the ? character in the filter as shown in the figure below.
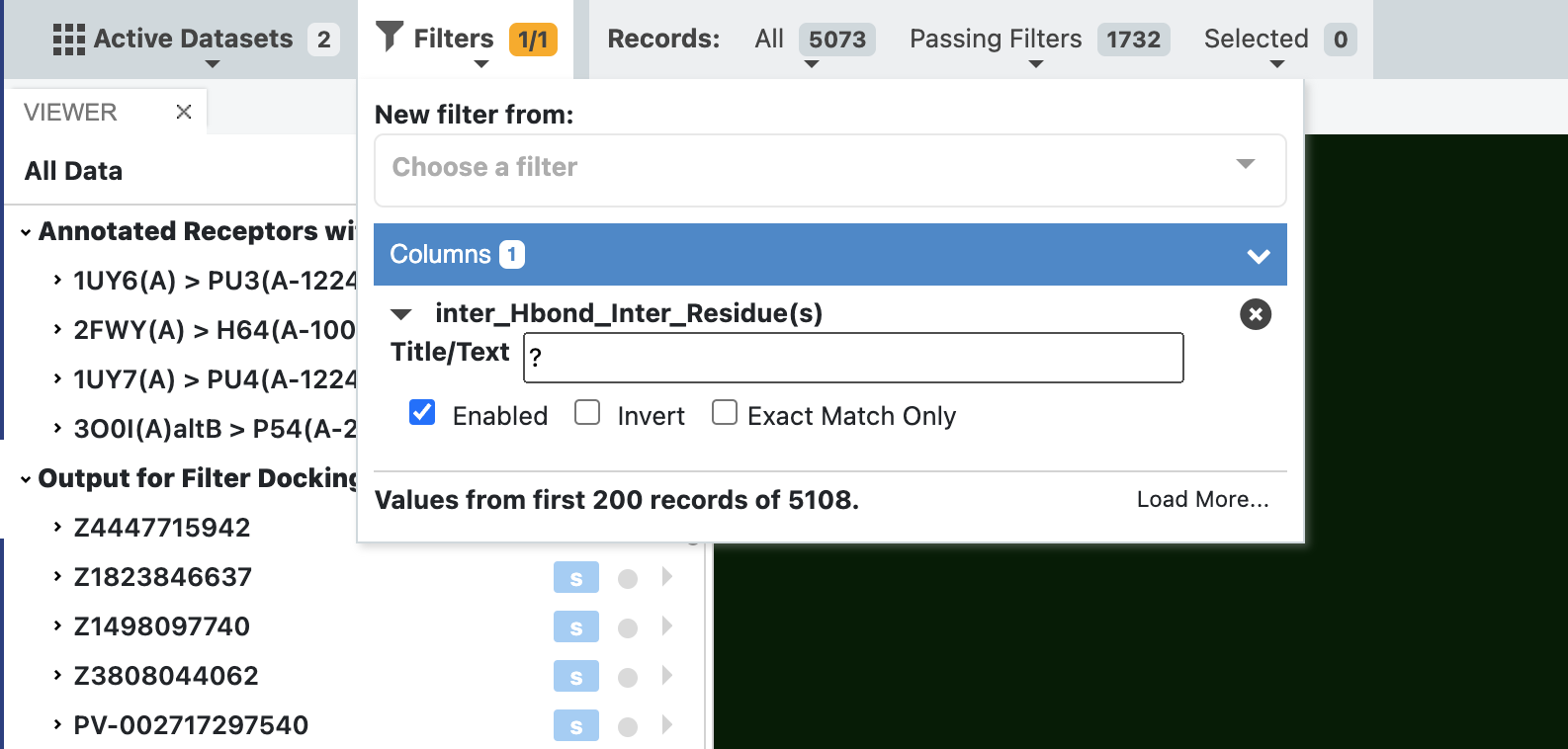
Filter dataset for a given interaction using a wildcard¶
Once filtering has been applied, the number of records passing the filter is reduced to 1732 from the input 5073 records (all). We can now step through these poses and see the results in the 3D viewer, with interactions being drawn in 3D.
Note
There are instances where an interaction will not be drawn even if the filter states that an interaction is present. This is because during interaction perception, the hydrogen bond network was re-optimized for each target-ligand pair. These results were not stored for visualization, but only retained for annotations for filtering. In these cases it is likely very intuitive to see that a Histidine or Asparagine residue could flip or change state to accommodate a hydrogen bond. This flipping or state changing is done implicitly by the docking algorithm; hence we are annotating the interaction.
Note that datasets from GigaDocking contain a field called “Design Unit ID”, indicating which receptor a given molecule was docked against. Use this field along with the same field on the receptor dataset to correctly display the target-ligand poses and have appropriate interactions drawn.
In the above example we identified all poses with a hydrogen bond between the docked molecule and the receptor, but in a lot of cases, it is interesting to explore which molecules form a hydrogen bond (or any other type of interaction) with a specific residue. In this case we can change the filter to only pass docked poses that have an interaction with a specific residue. This is illustrated in the figure below, where it can also be seen that there are only 484 poses with this specific interaction.
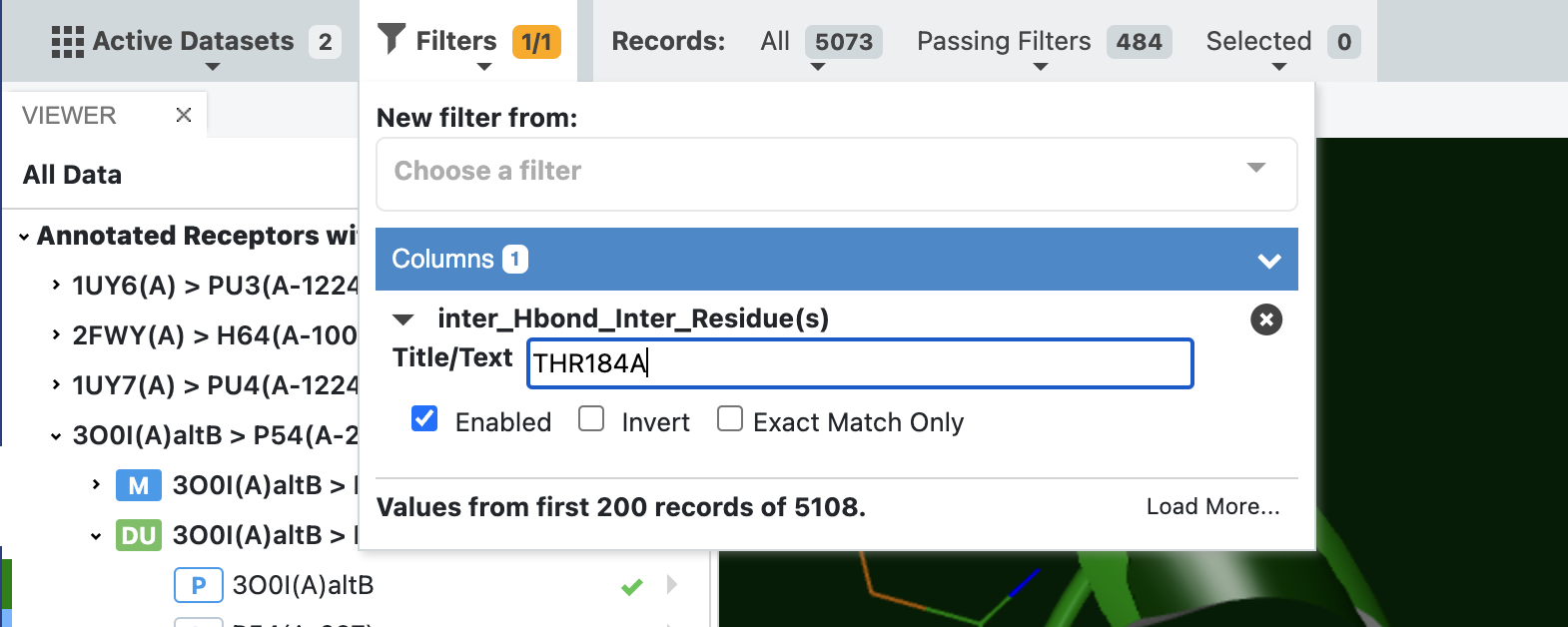
Filter dataset by a specific residue for a given interaction¶
It is also possible to filter for two specific hydrogen bonds by applying the same filter twice with a specific residue each. Multiple filters can be applied at the same time, but that is beyond the scope of this tutorial.
Next steps¶
These triaged subset of datasets can be saved as new datasets and carried forward for further inspection, e.g. Short Trajectory Molecular Dynamics (STMD).