Substructure Search with MDL Queries
Section Substructure Search describes how to perform substructure
search initialized with a SMILES or SMARTS string.
OEChem TK also provides the ability to interpret and utilize query structures
expressed in the MDL query file format
(see MDL query example in Figure: Example of MDL query substructure).
Listing 1 shows how to initialize an
OESubSearch object from a MDL query file and
perform a substructure search.
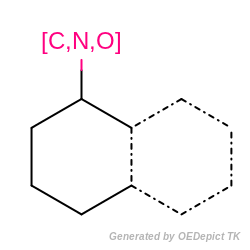
Example of MDL query substructure
Listing 1: Example of substructure search using MDL query file
from openeye import oechem
qfile = oechem.oemolistream("query.mol")
tfile = oechem.oemolistream("targets.sdf")
# set the same aromaticity model for the query and the target file
aromodel = oechem.OEIFlavor_Generic_OEAroModelMDL
qflavor = qfile.GetFlavor(qfile.GetFormat())
qfile.SetFlavor(qfile.GetFormat(), (qflavor | aromodel))
tflavor = tfile.GetFlavor(tfile.GetFormat())
tfile.SetFlavor(tfile.GetFormat(), (tflavor | aromodel))
# read MDL query and initialize the substructure search
opts = oechem.OEMDLQueryOpts_Default | oechem.OEMDLQueryOpts_SuppressExplicitH
qmol = oechem.OEQMol()
oechem.OEReadMDLQueryFile(qfile, qmol, opts)
ss = oechem.OESubSearch(qmol)
# loop over target structures
tindex = 1
for tmol in tfile.GetOEGraphMols():
oechem.OEPrepareSearch(tmol, ss)
if ss.SingleMatch(tmol):
print("hit target =", tindex)
tindex += 1
After opening the MDL query and the target files, the model used to assign aromaticity to the imported structures can be adjusted.
aromodel = oechem.OEIFlavor_Generic_OEAroModelMDL
qflavor = qfile.GetFlavor(qfile.GetFormat())
qfile.SetFlavor(qfile.GetFormat(), (qflavor | aromodel))
tflavor = tfile.GetFlavor(tfile.GetFormat())
tfile.SetFlavor(tfile.GetFormat(), (tflavor | aromodel))
If the aromaticity model is not specified for the input files, then the OpenEye’s aromaticity model is used by default. For more information about the various aromaticity models of OEChem TK see Aromaticity Perception.
oechem.OEReadMDLQueryFile(qfile, qmol, opts)
ss = oechem.OESubSearch(qmol)
In general, the aromaticity model chosen should be consistent between the query and target molecules to be searched. Using different aromaticity models may produce false negatives as aromatic systems may be treated differently. Section Aromaticity further explains the effects of using various aromaticity models when performing a substructure search.
OEReadMDLQueryFile function reads the
MDL query directly into a OEQMolBase object ,
which then can be used to initialize an OESubSearch
instance.
The MDL query structure can also be read into a OEMolBase
object (see code snippet below).
In this case, the OEReadMDLQueryFile
function attaches the query features present in the input MDL file to the related
atoms and bonds of the OEMolBase object.
The OEQMolBase object can be subsequently created by
calling the OEBuildMDLQueryExpressions
function.
mol = OEGraphMol()
OEReadMDLQueryFile(qfile,mol)
# mol can be manipulated here
qmol = OEQMol()
# build OEQMol with OEMDLQueryOpts_Default option
OEBuildMDLQueryExpressions(qmol,mol)
ss = OESubSearch(qmol)
The declaration of these functions are:
OEReadMDLQueryFile(ifs,mol)
# ifs-oemolistream, mol-OEMolBase
# returns true or false
OEReadMDLQueryFile(ifs,qmol,opts)
# ifs-oemolistream, mol-OEQMolBase, opts-integer with OEMDLQueryOpts_Default
# returns true or false
OEBuildMDLQueryExpressions(qmol,mol,opts)
# ifs-oemolistream, mol-OEQMolBase, opts-integer with OEMDLQueryOpts_Default
# returns true or false
MDL Query Interpretation
The opts parameter defines how the MDL query is interpreted when
an OEQMolBase object is constructed.
The following options are present in the OEMDLQueryOpts
namespace:
-
Only constraints explicitly specified in the MDL file are added to the OEQMolBase query structure. See section Supported MDL Query Features for the supported MDL query features.
OEMDLQueryOpts_SuppressExplicitHThis option controls how the explicit hydrogens of the query are matched to the explicit/implicit hydrogens of the target structures. For more information see Explicit Hydrogens.
OEMDLQueryOpts_AddBondAliphaticConstraintIf this option is specified, then an aliphatic query bond can only be mapped to the aliphatic bonds in the target structure. Figure: Interpretation A shows how the MDL query structure is interpreted when the
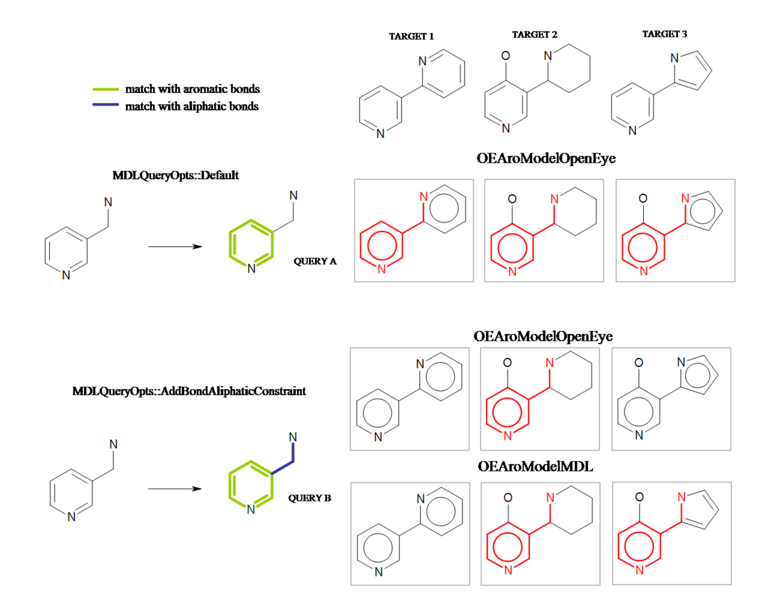
OEMDLQueryOpts_AddBondAliphaticConstraintoption is used.Query ‘A’ will match all three of the target compounds displayed in Figure: Interpretation A. If the OpenEye model is used to perceive aromatic rings, then query ‘B’ substructure is present only in target ‘2’. If the MDL aromaticity model is used, then target ‘3’ is also a hit, since five-membered heterocycles are not considered aromatic in this model. For more information about different aromaticity models and their effects on substructure searching see section Aromaticity.
Interpretation A
Interpretation of the MDL query with ‘AddBondAliphaticConstraint’ option.
OEMDLQueryOpts_AddBondTopologyConstraintBy default, a bond that is part of any ring system in the query structure can only be mapped to ring bonds in the target structure. If the
OEMDLQueryOpts_AddBondTopologyConstraintoption is specified, constraints are also added to the chain bonds of the query in order to map them to only chain bonds in the target.Figure: Interpretation B shows how the MDL query structure is interpreted when the
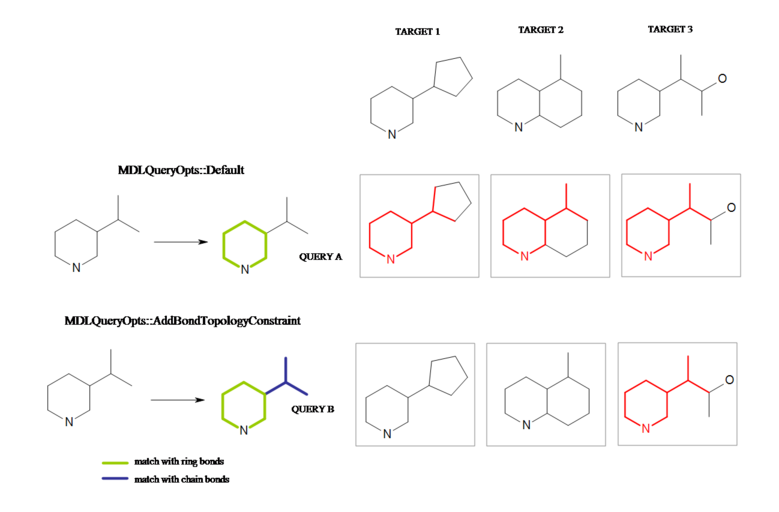
OEMDLQueryOpts_AddBondTopologyConstraintoption is used. Query ‘A’ will match all three of the target compounds displayed in Figure: Interpretation B, while query ‘B’ is present only in target ‘3’.If the ‘ring’ property is specified in the MDL query file for a particular chain bond in the query structure, then its topology constraint is overridden.
Interpretation B
Interpretation of the MDL query with `AddBondTopologyConstrain’ option.
OEMDLQueryOpts_MatchAtomStereoWhen this option is enabled, an S/R atom stereo configuration in the query will only match any S/R configuration in the target molecule, but not to a R/S (opposite) or an unspecified one. If an atom stereo configuration is undefined in the query, it is allowed to match to any target atom regardless of its stereo configuration.
-
When this option is enabled, query atoms with an isotope can match to a target atom only if they have the same atomic mass. If the query atom has no specified isotope number, it will match to any target atom regardless of its atomic mass.
Supported MDL Query Features
The table below summarizes the MDL query features currently supported by the OEChem TK.
These query feature can only be read with the low level OEReadMDLQueryFile
function and not the high-level OEReadMolecule function.
CTFile V2K |
CTFile V3K |
Notes |
|---|---|---|
|
|
formal charge |
|
|
number of allowed hydrogens V2K:0=off, 1=H0, 2-5 for H1-H4; V3K:0=off, -1=H0, 1-5 for H1-H5 |
|
|
mass difference |
|
|
limit of number of allowed ring bonds attached to an atom |
|
|
number of allowed substitutions of an atom |
|
|
list alternative atom types for an atom |
|
|
stereo care (e.g. for c/t doubles) |
|
|
L=atom list, A=Any atom except hydrogen, Q=Any atom except hydrogen and carbon |
|
|
specifies whether or not an atom is unsaturated, i.e., having at least one multiple bond |
|
|
bond stereochemistry that is considered, if both ends of the bond are marked with stereo care flags in the atom block |
|
|
bond topology (0=off, 1=ring, 2=chain) |
|
|
4-aromatic, 5=single or double, 6=single or aromatic, 7=double or aromatic, 8=any |
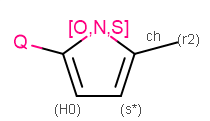
Query
Example of MDL query structure.
See also
MDL Query Depiction chapter for the depiction of the supported query features
Accelrys CTfile Formats document for more information about the atom and bond query features
Aromaticity
Perceiving aromaticity in the query and the target structures is important in order to ensure that the result of a substructure search is independent of the different Kekulé representations of the participating structures. A query bond which is part of an aromatic ring system can be mapped to any aromatic bonds in the target. Figure: Aromaticity Match shows an example where both Kekulé representations of the benzene-1,2-diol substructure are present in the two target structures.
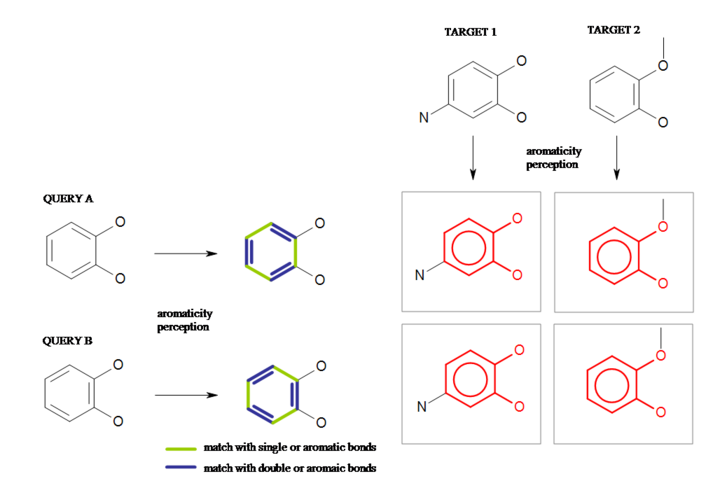
Aromaticity Match
The result of substructure search is independent of the Kekulé representations of the participating structures.
Altering the aromaticity model will affect the results of a substructure search. Figure: Aromaticity A – Figure: Aromaticity C show several examples where different results were obtained by applying the MDL and the OpenEye aromaticity models. This is a consequence of the fact that in the MDL aromaticity model, five-membered heterocycles are not considered aromatic.
Note
It is highly recommended to apply the same aromaticity model to the query and the target structures.
Listing 1 shows an example of how to
change the aromaticity flavor of input files.
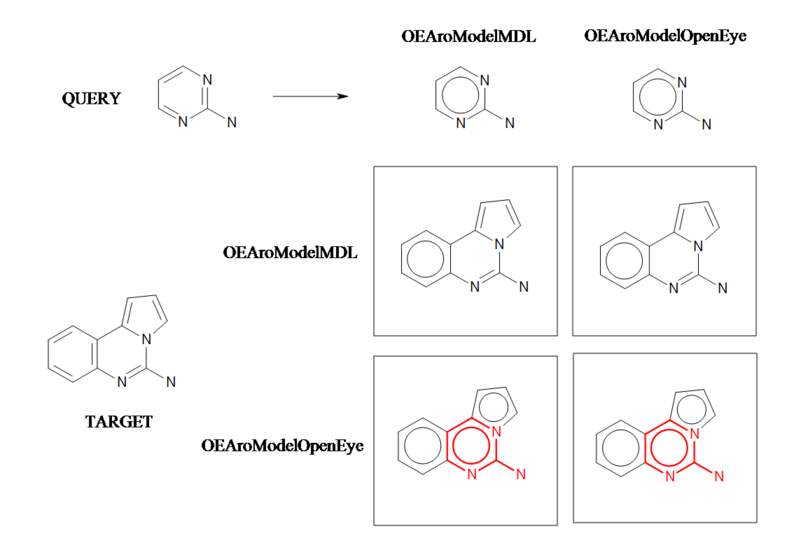
Aromaticity A
Example of a substructure search applying different aromaticity models.
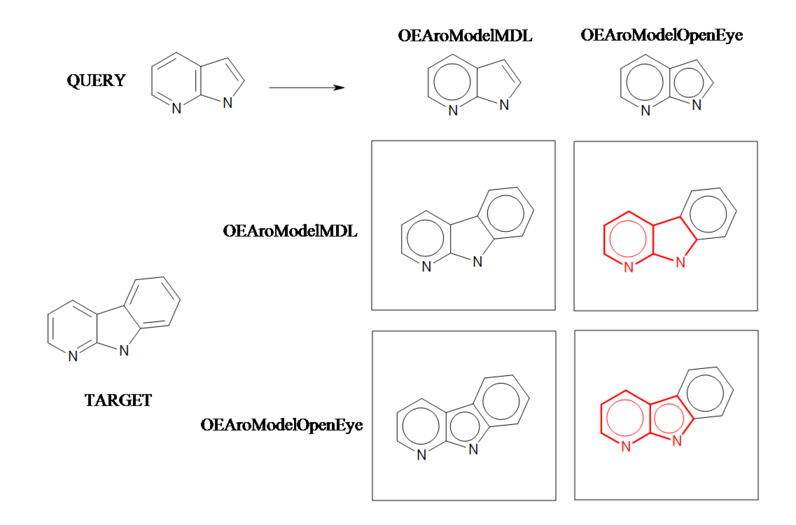
Aromaticity B
Example of a substructure search applying different aromaticity models.
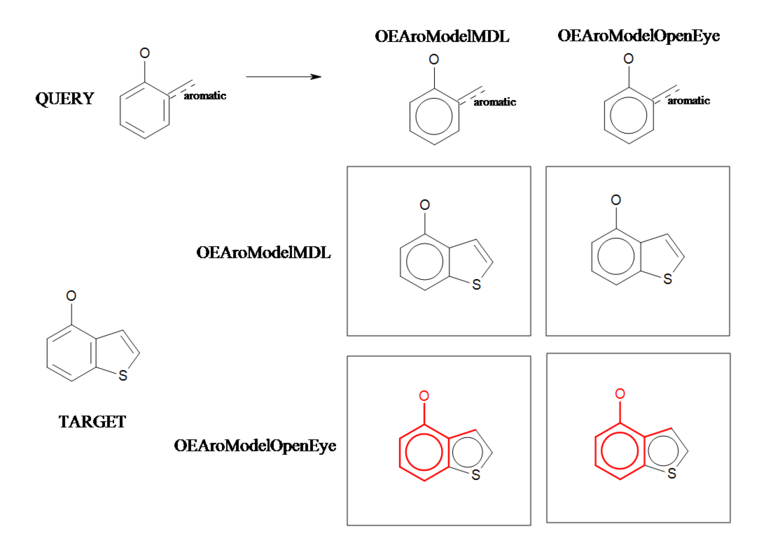
Aromaticity C
Example of a substructure search applying different aromaticity models.
Aromaticity with Generic Atoms
If a ring in the query structure contains generic atom(s) (see example in Figure: Generic atom example A), then the aromaticity of the ring can not be perceived. In order to maintain the independence from the Kekulé representation, 6-membered rings with alternating single/double bonds are assumed to be aromatic.
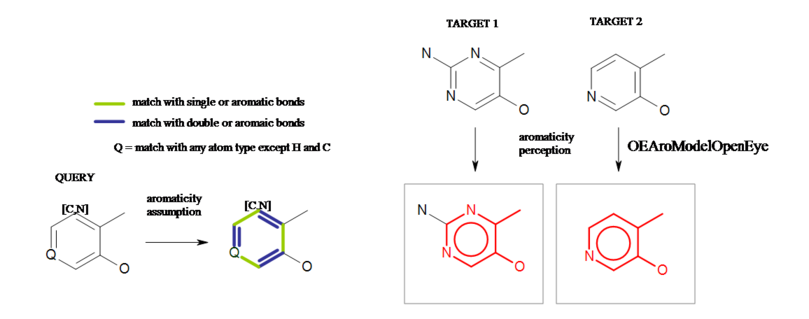
Generic atom example A
Example of aromaticity assumption in a 6-membered ring with generic atoms.
Similarly, a 5-membered ring with generic atom(s) is considered aromatic if it is composed of two single and two double bonds. See Example in Figure: Generic atom example B.
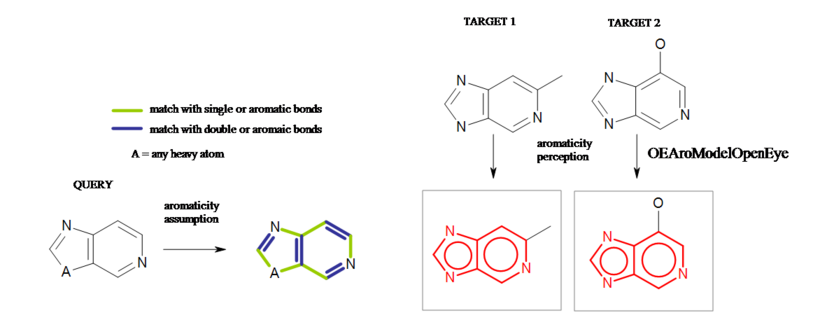
Generic atom example B
Example of aromaticity assumption in a 5-membered ring with generic atoms.
Explicit Hydrogens
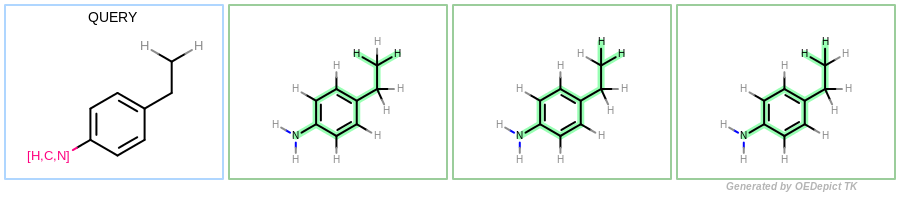
During the substructure search, each query atom has to be mapped to a target atom in order to detect subgraph isomorphism. Therefore, a problem can arise if the query structure contains explicit hydrogens or an atom list with hydrogens (see example in Figure: Query), but the target structure has implicit hydrogens Figure: Targets)
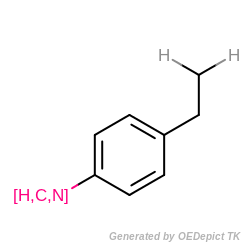
Example of MDL query structure with explicit hydrogens
Listing 2: Example of substructure search with accessing atom mapping
from openeye import oechem
qfile = oechem.oemolistream("query.mol")
tfile = oechem.oemolistream("targets.sdf")
# read MDL query and initialize the substructure search
qmol = oechem.OEQMol()
oechem.OEReadMDLQueryFile(qfile, qmol)
ss = oechem.OESubSearch(qmol)
# loop over target structures
tindex = 1
for tmol in tfile.GetOEGraphMols():
oechem.OEAddExplicitHydrogens(tmol)
oechem.OEPrepareSearch(tmol, ss)
for mi in ss.Match(tmol, True):
print("hit target = %d" % tindex, end=" ")
for ai in mi.GetTargetAtoms():
print("%d%s" % (ai.GetIdx(), oechem.OEGetAtomicSymbol(ai.GetAtomicNum())), end=" ")
print()
tindex += 1
This problem can be solved in two ways:
The explicit hydrogens in the query molecule can be suppressed during
OEQMolBaseconstruction by using theOEMDLQueryOpts_SuppressExplicitHoption (see example inListing 1). A query atom can only be mapped to a target atom if it has at least as many implicit hydrogens as explicit hydrogens were removed from the query atom. This solution is recommended only if the presence or absence of the query substructure is of interest, but not the complete mapping between the query and the target. TheSingleMatchfunction returns whether or not the query is present in the target, but the mapping is not accessible.
If the complete mapping between the query and the target is of interest, the
OEAddExplicitHydrogensfunction has to be called for each target structure before performing the substructure search. (See example inListing 2).
In both cases, the query presented in Figure: Query will match only target ‘C’ and ‘D’ shown in Figure: Targets. Figure: Atom mapping shows the three detected substructures in target ‘C’ when adding explicit hydrogens to the target structures.
The execution of the substructure search is significantly faster if the hydrogens are suppressed in the target structures, since the search space can be an order of magnitude smaller.
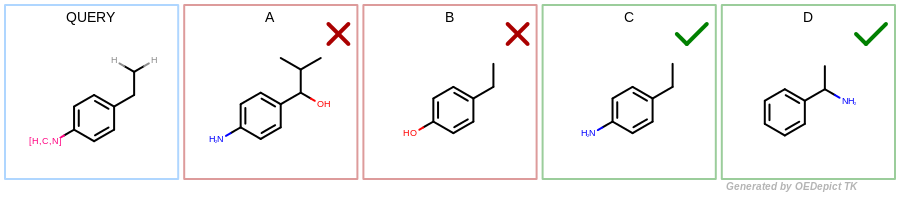
Targets – Example of target structures with implicit hydrogens
Atom mapping – Example of substructure search with atom mapping