OEGetRingLinkerSideChainFragments
OESystem::OEIterBase<OEChem::OEAtomBondSet>* OEGetRingLinkerSideChainFragments(
const OEChem::OEMolBase& mol,
const OEBemisMurckoOptions& options = OEBemisMurckoOptions());
Warning
The following function is deprecated. Please use the more generic function above.
OESystem::OEIterBase<OEChem::OEAtomBondSet>* OEGetRingLinkerSideChainFragments(
const OEChem::OEMolBase& mol,
unsigned perceiveType)
The OEGetRingLinkerSideChainFragments function partitions
the given molecule into ring, linker and side chain fragments as defined
in [Bemis-1996] .
The OEBemisMurckoOptions class can be used to
specify both the OERegionType and also add additional
functional groups to fragments. The perceiveType parameter only allows modification
of the OERegionType of the fragments returned.
It returns an iterator over OEAtomBondSet objects where
each OEAtomBondSet container stores the atoms and the bonds
of an identified fragment.
See example in Figure: Example of fragments returned by the
OEGetRingLinkerSideChainFragments function
where each fragment returned by the OEGetRingLinkerSideChainFragments
function is highlighted with a different color.
The perceived regions are based upon the heavy atoms in the molecule. If the molecule contains explicit hydrogen atoms, those explicit hydrogens will be returned in the same perception set(s) as their alpha heavy atom neighbor. If the perception set(s) returned are used to subset or fragment the molecule, a mixture of implicit and explicit hydrogens may result, so care should be taken to normalize the structure for subsequent use.
Definitions from [Bemis-1996]:
- ring systems
Cycles within the graph representation of molecules and cycles sharing an edge (a connection between two atoms or a bond).
- linker atoms
Atoms that are on the direct path connecting two ring systems are defined as linker atoms.
- side chain atoms
Any nonring, nonlinker atoms are defined as side chain atoms.
Note
The molecule argument is const, so this function requires that the
OEFindRingAtomsAndBonds perception activity has been
performed prior to calling this function.
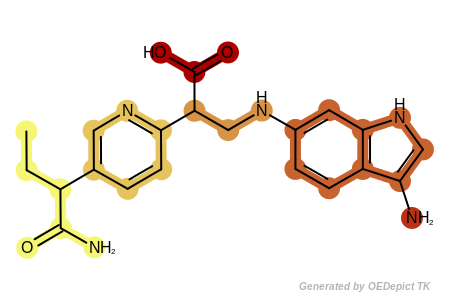
Example of fragments returned by the OEGetRingLinkerSideChainFragments function
See also
OEAtomBondSet and OEMolBase classes in the OEChem TK manual
Molecule Fragmentation chapter