OEHermiteShapeFunc
class OESHAPE_API OEHermiteShapeFunc : public OEShapeFunc
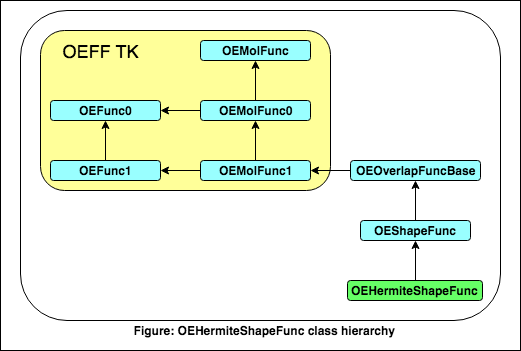
This class is derived from OEShapeFunc and is used to approximate Tanimoto shape-similarity coefficient based on Hermite representation of molecules.
See also
OEHermite class
OEHermiteOptions class
Shape from Hermite expansion examples
- The following methods are publicly inherited from OEFunc0:
- The following methods are publicly inherited from OEFunc1:
- The following methods are publicly inherited from OEMolFunc:
- The following methods are publicly inherited from OEOverlapFuncBase:
- The following methods are publicly inherited from OEShapeFunc:
- The OEHermiteShapeFunc class defines the following public methods:
Constructors
OEHermiteShapeFunc(const OEHermiteShapeFunc &)
OEHermiteShapeFunc(const OEShapeOptions &argOptions=OEShapeOptions())
OEHermiteShapeFunc(const OEHermiteOptions &HermOptRef,
const OEHermiteOptions &HermOptFit)
OEHermiteShapeFunc(const OEShapeOptions &argOptions,
const OEHermiteOptions &HermOptRef,
const OEHermiteOptions &HermOptFit)
The four constructors are in the respective order: a standard copy constructor, a constructor that inputs a OEShapeOptions object, a constructor that inputs two OEHermiteOptions objects, one for reference and one for fit molecules, and finally a constructor that inputs OEShapeOptions object along with two OEHermiteOptions objects, one for reference and one for fit molecules.
operator=
OEHermiteShapeFunc &operator=(const OEHermiteShapeFunc &)
The assignment operator.
GetFitHermiteOptions
const OEHermiteOptions &GetFitHermiteOptions() const
This method returns OEHermiteOptions class currently being used for the fit molecule.
GetRefHermiteOptions
const OEHermiteOptions &GetRefHermiteOptions() const
This method returns OEHermiteOptions class currently being used for the reference molecule.