Ring Perception
Problem
You want to determine whether two atoms belong to the same ring. See examples in Table 1.
Ring perception is one of the fundamental algorithms when handling chemical structures. While OEChem TK does not provide a solution for SSSR (Smallest Set of Smallest Rings) it includes wide range of functions (see Table 2) that answer related questions. This recipe will illustrate how to utilize these functions to determine whether two atoms belong to the same ring.
See also
Smallest Set of Smallest Rings (SSSR) Considered Harmful section in OEChem TK manual
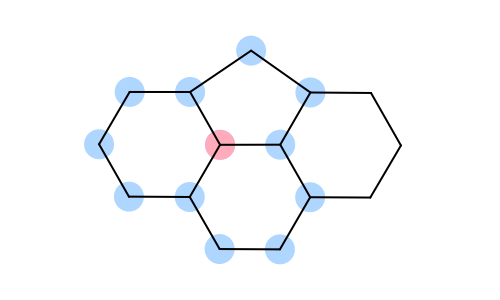
|
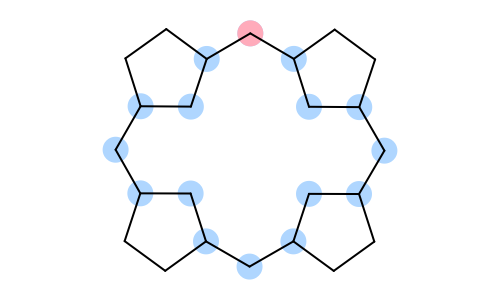
|
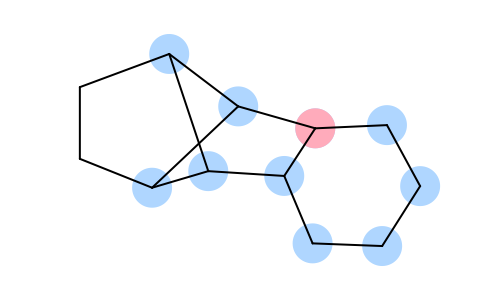
|
API |
See subsection |
|---|---|
Ring Perception section in OEChem TK |
|
Ingredients
|
Difficulty level


Download
Solution
The algorithm implemented in AtomsInSameRing is based on a simple concept: if two atoms belong to the same ring then there has to at least two alternative ring paths between them. These paths are identified by using the OEShortestPath function.
1def AtomsInSameRing(atomA, atomB):
2
3 if not atomA.IsInRing() or not atomB.IsInRing():
4 return False
5
6 if atomA == atomB:
7 return True
8
9 firstpath = [a for a in oechem.OEShortestPath(atomA, atomB, oechem.OEAtomIsInChain())]
10 firstpathlength = len(firstpath)
11
12 if firstpathlength == 2:
13 return True # neighbors
14
15 if firstpathlength == 0:
16 return False # not is same ring system
17
18 smallestA = oechem.OEAtomGetSmallestRingSize(atomA)
19 smallestB = oechem.OEAtomGetSmallestRingSize(atomB)
20
21 if firstpathlength > smallestA and firstpathlength > smallestB:
22 return False # too far away
23
24 # try to find the second shortest different path
25 excludepred = ChainAtomOrAlreadyTraversed(firstpath[1:-1])
26 secondpath = [a for a in oechem.OEShortestPath(atomA, atomB, excludepred)]
27 secondpathlength = len(secondpath)
28
29 if secondpathlength == 0:
30 return False # can not be in the same ring
31
32 if secondpathlength > smallestA and secondpathlength > smallestB:
33 return False # too far away
34
35 sumringsize = len(firstpath) + len(secondpath) - 2
36 if sumringsize > smallestA and sumringsize > smallestB:
37 return False
38
39 inringA = oechem.OEAtomIsInRingSize(atomA, sumringsize)
40 inringB = oechem.OEAtomIsInRingSize(atomB, sumringsize)
41 return inringA and inringB
ChainAtomOrAlreadyTraversed is the atom predicate that is used with OEShortestPath to identify a second alternative path between two atoms.
1class ChainAtomOrAlreadyTraversed(oechem.OEUnaryAtomPred):
2 def __init__(self, exclude):
3 oechem.OEUnaryAtomPred.__init__(self)
4 self.exclude = exclude
5
6 def __call__(self, atom):
7 if not atom.IsInRing():
8 return False
9 return (atom in self.exclude)
10
11 def CreateCopy(self):
12 return ChainAtomOrAlreadyTraversed(self.exclude).__disown__()
Usage
Usage
prompt > python3 samering.py .ism
c1ccc2c(c1)cc[nH]2
atom 0 C in the same ring as 0 C 1 C 2 C 3 C 4 C 5 C
atom 1 C in the same ring as 0 C 1 C 2 C 3 C 4 C 5 C
atom 2 C in the same ring as 0 C 1 C 2 C 3 C 4 C 5 C
atom 3 C in the same ring as 0 C 1 C 2 C 3 C 4 C 5 C 6 C 7 C 8 N
atom 4 C in the same ring as 0 C 1 C 2 C 3 C 4 C 5 C 6 C 7 C 8 N
atom 5 C in the same ring as 0 C 1 C 2 C 3 C 4 C 5 C
atom 6 C in the same ring as 3 C 4 C 6 C 7 C 8 N
atom 7 C in the same ring as 3 C 4 C 6 C 7 C 8 N
atom 8 N in the same ring as 3 C 4 C 6 C 7 C 8 N
Discussion
The following examples show how to use ring perception functions available in OEChem TK.
Identifying Ring Systems of a Molecule
nrrings, ringlist = oechem.OEDetermineRingSystems(mol)
disp = oedepict.OE2DMolDisplay(mol, opts)
highlight = oedepict.OEHighlightByLasso(oechem.OEBlack)
highlight.SetConsiderAtomLabelBoundingBox(True)
ringpred = oechem.OEPartPredAtom(ringlist)
for ringidx, color in zip(range(1, nrrings + 1), oechem.OEGetVividColors()):
ringpred.SelectPart(ringidx)
ringset = oechem.OEAtomBondSet(mol.GetAtoms(ringpred))
highlight.SetColor(color)
oedepict.OEAddHighlighting(disp, highlight, ringset)
oedepict.OERenderMolecule(image, disp)
Download code |
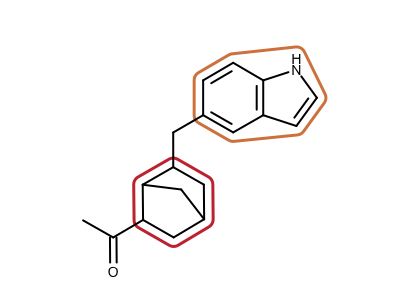
|
See also
OEDetermineRingSystems method
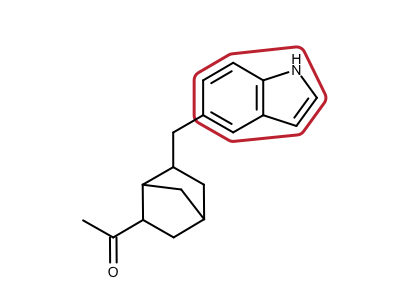
Identifying Aromatic Ring Systems of a Molecule
nrrings, ringlist = oechem.OEDetermineAromaticRingSystems(mol)
disp = oedepict.OE2DMolDisplay(mol, opts)
highlight = oedepict.OEHighlightByLasso(oechem.OEBlack)
highlight.SetConsiderAtomLabelBoundingBox(True)
ringpred = oechem.OEPartPredAtom(ringlist)
for ringidx, color in zip(range(1, nrrings + 1), oechem.OEGetVividColors()):
ringpred.SelectPart(ringidx)
ringset = oechem.OEAtomBondSet(mol.GetAtoms(ringpred))
highlight.SetColor(color)
oedepict.OEAddHighlighting(disp, highlight, ringset)
oedepict.OERenderMolecule(image, disp)
Download code |
|
See also
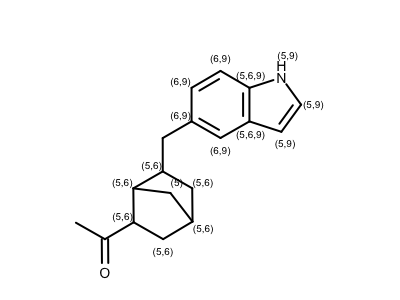
Identifying Atom in Certain Ring Size
class LabelRingSize(oedepict.OEDisplayAtomPropBase):
def __init__(self, maxringsize):
self.maxringsize = maxringsize
oedepict.OEDisplayAtomPropBase.__init__(self)
def __call__(self, atom):
if not atom.IsInRing():
return ""
rings = []
for ringsize in range(3, self.maxringsize):
if oechem.OEAtomIsInRingSize(atom, ringsize):
rings.append(ringsize)
if len(rings) == 0:
return ""
return "(" + ",".join([str(r) for r in rings]) + ")"
def CreateCopy(self):
return LabelRingSize(self.maxringsize).__disown__()
scale = oedepict.OEScale_AutoScale
opts = oedepict.OE2DMolDisplayOptions(width, height, scale)
opts.SetAtomColorStyle(oedepict.OEAtomColorStyle_WhiteMonochrome)
opts.SetAtomPropertyFunctor(LabelRingSize(maxringsize=10))
disp = oedepict.OE2DMolDisplay(mol, opts)
Download code |
|
See also
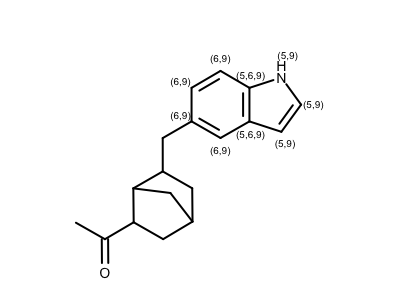
Identifying Atom in Certain Aromatic Ring Size
class LabelAromaticRingSize(oedepict.OEDisplayAtomPropBase):
def __init__(self, maxringsize):
self.maxringsize = maxringsize
oedepict.OEDisplayAtomPropBase.__init__(self)
def __call__(self, atom):
if not atom.IsInRing():
return ""
rings = []
for ringsize in range(3, self.maxringsize):
if oechem.OEAtomIsInAromaticRingSize(atom, ringsize):
rings.append(ringsize)
if len(rings) == 0:
return ""
return "(" + ",".join([str(r) for r in rings]) + ")"
def CreateCopy(self):
return LabelAromaticRingSize(self.maxringsize).__disown__()
scale = oedepict.OEScale_AutoScale
opts = oedepict.OE2DMolDisplayOptions(width, height, scale)
opts.SetAtomColorStyle(oedepict.OEAtomColorStyle_WhiteMonochrome)
opts.SetAtomPropertyFunctor(LabelAromaticRingSize(maxringsize=10))
disp = oedepict.OE2DMolDisplay(mol, opts)
Download code |
|
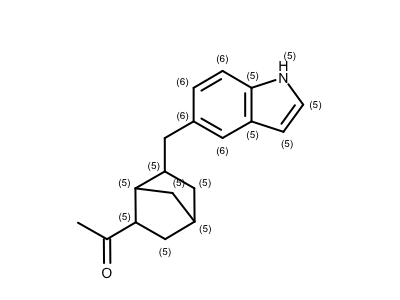
Identifying Atoms’ Smallest Ring Size
class LabelSmallestRingSize(oedepict.OEDisplayAtomPropBase):
def __init__(self):
oedepict.OEDisplayAtomPropBase.__init__(self)
def __call__(self, atom):
if not atom.IsInRing():
return ""
smallest = oechem.OEAtomGetSmallestRingSize(atom)
return "(" + str(smallest) + ")"
def CreateCopy(self):
return LabelSmallestRingSize().__disown__()
scale = oedepict.OEScale_AutoScale
opts = oedepict.OE2DMolDisplayOptions(width, height, scale)
opts.SetAtomColorStyle(oedepict.OEAtomColorStyle_WhiteMonochrome)
opts.SetAtomPropertyFunctor(LabelSmallestRingSize())
disp = oedepict.OE2DMolDisplay(mol, opts)
Download code |
|
See also in OEChem TK manual
Theory
API
OEAtomBondSet class
OEPartPredAtom predicate