Depicting Molecular Graph Symmetry
Problem
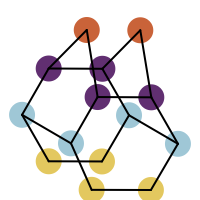
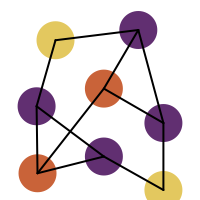
You want to easily identify whether there is a graph symmetry in a molecule. See examples in in Table 1.
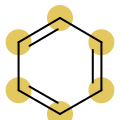
|
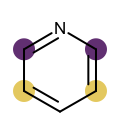
|
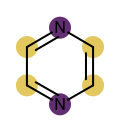
|
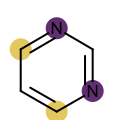
|
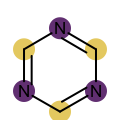
|
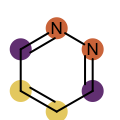
|
Ingredients
|
Difficulty Level

Solution
In DepictSymmetry function, the OEPerceiveSymmetry function is called to assign symmetry class for each atom of a given molecule. The symmetry class of each atom then can be retrieved using the OEAtomBase.GetSymmetryClass method.
After perceiving the molecule symmetry, atoms that belong to the same symmetry class are collected in a dictionary (lines 5-11). If a symmetry class only contains one atom, than it is considered unique and removed from the dictionary (lines 13-19). The atoms of the remaining symmetry classes are then highlighted with different colors (lines 24-26).
1def DepictSymmetry(image, mol, opts):
2
3 oechem.OEPerceiveSymmetry(mol)
4
5 symdict = dict()
6
7 for atom in mol.GetAtoms():
8 sym = atom.GetSymmetryClass()
9 if sym not in symdict:
10 symdict[sym] = oechem.OEAtomBondSet()
11 symdict[sym].AddAtom(atom)
12
13 uniquesym = []
14 for sym, atomset in symdict.items():
15 if atomset.NumAtoms() == 1:
16 uniquesym.append(sym)
17
18 for sym in uniquesym:
19 del symdict[sym]
20
21 oedepict.OEPrepareDepiction(mol)
22 disp = oedepict.OE2DMolDisplay(mol, opts)
23
24 colors = [c for c in oechem.OEGetContrastColors()]
25 for (sym, atomset), color in zip(symdict.items(), colors):
26 oedepict.OEAddHighlighting(disp, color, oedepict.OEHighlightStyle_BallAndStick, atomset)
27
28 oedepict.OERenderMolecule(image, disp)
Download code
Usage:
prompt > python3 symmetry2img.py
Discussion
Perceiving graph symmetry plays an important role in cheminformatics. The classification provided by the OEPerceiveSymmetry function can be used to:
identify chiral atoms
canonicalize molecules
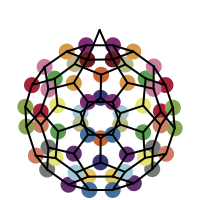
|
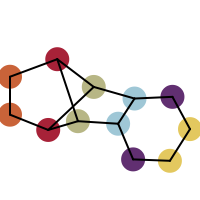
|
|
|
See also in OEChem TK manual
API
OEAtomBase.GetSymmetryClass method
OEAtomBondSet class
OEGetContrastColors function
OEPerceiveSymmetry function
See also in OEDepict TK manual
Theory
Highlighting chapter
API
OE2DMolDisplay class
OE2DMolDisplayOptions class
OEAddHighlighting function
OERenderMolecule function