FastROCS Plus with Docking and Consensus Score of Top Molecules
In this tutorial, a database collection provided by OpenEye will be searched with FastROCS for molecules similar to a ligand crystallized with HSP90. The top-scoring molecules from FastROCS will then be rescored using ROCS and docking into the HSP90 receptor and the generated consensus hit list will be examined. Running all the floes in this tutorial will cost approximately $30.
This tutorial uses the following floes:
Note
If you have not created a tutorial project or prepared the HSP90 heat shock protein 90 (HSP90) target (using crystal structure 1uyg) to create the hsp90_design_unit dataset, please go to the Preliminary Setup Tutorial before beginning this tutorial.
Run the FastROCS Plus Floe
Locate the FastROCS Plus Floe on the Workflows tab in the Floe page.
Click “Launch Floe” to bring up the Job Form and set the following parameters.
Output Path: Use Tutorial/My Data/FastROCS HSP90 Consensus as the path.
Inputs
Input Query Dataset(s): Click “Choose Input” and select the hsp90_design_unit dataset from the SPRUCE - Protein Preparation Floe.
Note
This floe automatically uses the bound ligand when a design unit with a ligand is passed to this parameter. Outside of the tutorial, you might alternatively pass one or more known active molecules to this parameter.
FastROCS Input Collection: Follow the path Organization Data/OpenEye Data/FastROCS Collections and select an appropriate collection for your needs. To run the most affordable example for this tutorial, select the smallest collection.
Design Unit(s) (Optional): Select the hsp90_design_unit dataset.
Click “Start Job” to begin the floe. The job will take roughly 30 minutes to run and cost about $30.
Note
This floe produces the following seven hit lists, although you will only use three in this tutorial.
FastROCS Hit List: Output dataset that contains the top hits directly from FastROCS.
ROCS Hit List: Output hit list dataset from ROCS rescoring of the top FastROCS hits.
Dock Hit List: Output hit list dataset from docking the top FastROCS hits.
Consensus ROCS Hit List: Consensus output hit list ranked by ROCS Combo Tanimoto.
Consensus Dock Hit List: Consensus output hit list ranked by docking score.
FastROCS Novelty Hit List: Output dataset that contains a subset of top hits from FastROCS that are highly diverse in 2D.
ROCS Novelty Hit List: Output hit list dataset that contains a subset of top hits from ROCS rescoring that are highly diverse in 2D.
View Consensus Result
Activate the Consensus Dock, Dock, and ROCS Hit Lists
Navigate to the Data page from the blue navigation bar.
In the My Data folder, select the FastROCS HSP90 Consensus subfolder.
In the ‘Show’ drop-down menu, be sure that the Datasets option is selected.
Make the following datasets active by clicking on the white circle with the plus symbol which will then become a green checkmark.
Consensus Dock Hit List
ROCS Hit List
Dock Hit List
These datasets should appear in the ‘Active Datasets’ drop-down. If other datasets are active, you can deselect them in the list or click the “Clear All” button.
For more detailed information on how to navigate the 3D & Analyze page, and in particular the 3D Viewer, please see the User Guide.
Next, switch to the 3D & Analyze page and select the 3D Analyze layout.
Plot the Dock and ROCS Scores in the 3D & Analyze Page and Highlight the Consensus Result
In the plot, set the axes for the docking and ROCS overlay scores:
Column X: Chemgauss4
Column Y: Tanimoto Combo
Note
Chemgauss4 is the docking score. Tanimoto Combo is the ROCS overlay score.
Once the plot has loaded, click the gear icon to open the Plot Options menu.
Color tab
Set the Color Dimension to Dataset Name.
Set a dark color for the Consensus Dock Hit List.
Set a light color for the Dock Hit List and the ROCS Hit List.
Marker tab
Set the Marker Dimension to Dataset Name.
Set the Consensus Dock Hit List marker to the plus sign.
Set the Dock Hit List and ROCS Hit List markers to open circles.
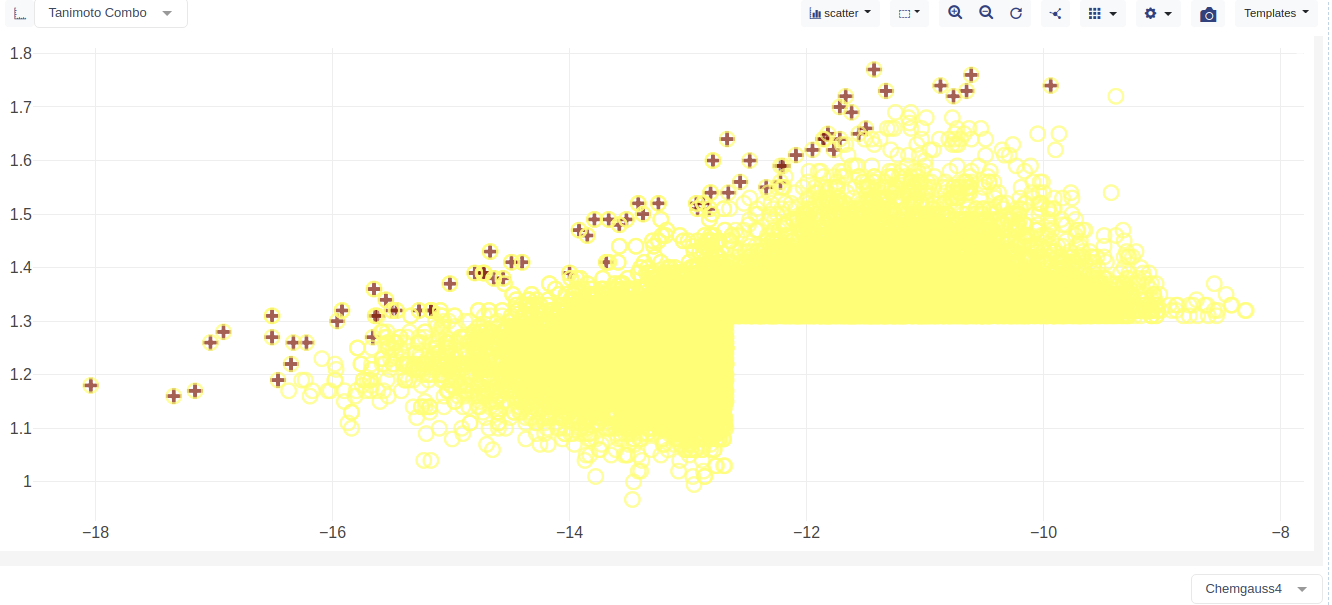
The plot will now show the plot of the hit list molecules dock score (i.e., Chemgauss4) versus the ROCS score (i.e., Tanimoto Combo). The consensus molecules are marked with a + symbol.
See also
Note
Some molecules appear in more than one hit list.
Observe the Consensus Docked Results in the 3D Viewer
On the Data page, navigate to the FastROCS HSP90 Consensus subfolder. and make the Design Units dataset active.
Navigate to the 3D & Analyze page and select the 3D Analyze view from the Layout drop-down. Click the X on the Plot panel to remove it from the page view.
In the top data tree, click the three dots in the bottom right and select Large Molecules.
Click the dot next to 1YUG(A) > PU2(A-1224) in the Large Molecule tree to log the design unit to the 3D view.
Open Data Handling modal from the Active Data Bar.
Click the All checkbox at the top of the All column. This will deselect all checkboxes in the All column.
Next, in the All column, check the boxes for Docked Molecule, Chemgauss4, and Tanimoto Combo.
Click “Apply” to apply the changes and close the Data Handling modal. The Data Handling changes may take a few minutes to update.
You can now view the consensus hit list molecules and their scores in the spreadsheet, while also examining the docked poses in the 3D Viewer.