Determine if Gigadock Will Give Good Results with a Given Receptor
Context
Docking, like all virtual screening tools, can work better on some systems than others. Before a full Gigadock Floe is performed, the performance of the run can be estimated if there are molecules known to be active against the available target.
Note
If there are several receptors available for the given target (e.g., receptors with different constraints or receptors using different conformations of the protein), it is recommended that this procedure be used on each receptor, and the receptor with the best enrichment be used for the full run.
Procedure
Use the Filter Collection Floe to create a collection of ~100,000 molecules randomly selected from the Gigadock collection to be used in the full Gigadock run. See the Dock One Million Molecules with the Gigadock Floe Tutorial for an example.
Generate conformers of the known active molecules using the OMEGA - 3D Conformer Ensemble Generation Floe using the default ‘Classic’ for the Conformer Generation Mode parameter.
Run the Gigadock Floe with the following parameters.
Output Path: Create a new appropriate folder.
Inputs
Design Unit or Receptor Dataset: Select the dataset with the receptor.
Input Conformer Collection: Set this to the collection created in Step 1.
Actives Dataset (Optional): Set this to the dataset of active conformers created in Step 2.
Once the Gigadock Floe finishes, an AUC dataset will be created which contains the AUC value for how well the active molecules scored in comparison to the 100K collection of molecules. Obviously, some of the 100K may be active, but it is reasonable to assume that the vast majority are inactive and treat it as a background decoy set.
The Floe Report for the Gigadock job will also contain a histogram showing how the active molecules scored in comparison to the background 100K collection of molecules.
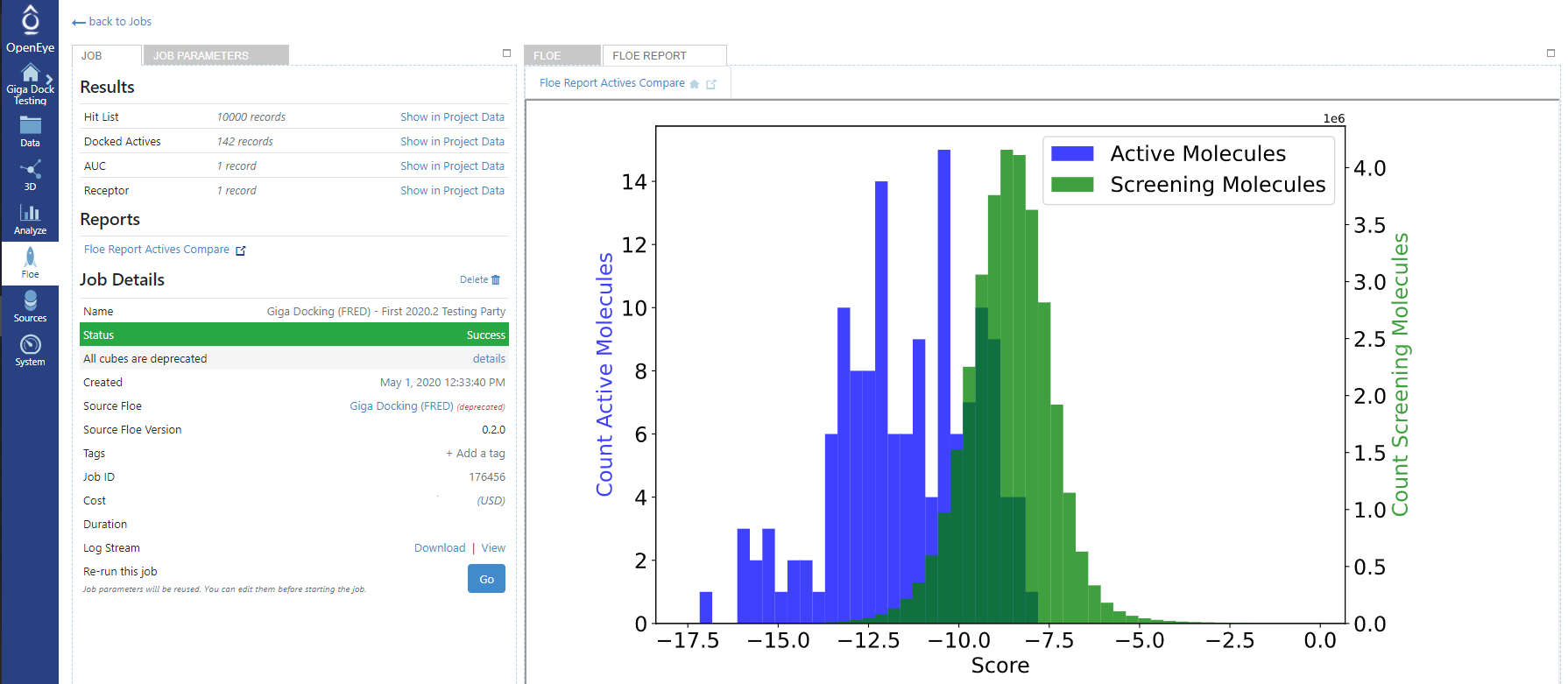
Figure 1. An example Floe Report for the Gigadock Floe.