Speed Up Preparation of a Very Large ROCS X 3D Library
Context
For very large libraries (>10 trillion products), preparing a 3D library can be slow and costly due to the bottleneck of generating conformers for the product sample. This is particularly an issue if the product samples tend to be large. For this type of library, it can be faster to skip conformer generation for the product sample and use an external hit list to seed the initial model.
Procedure
When preparing a 3D library with ROCS X - Prepare 3D Library, turn Enable Sample Products Off under Options: Sample Product.
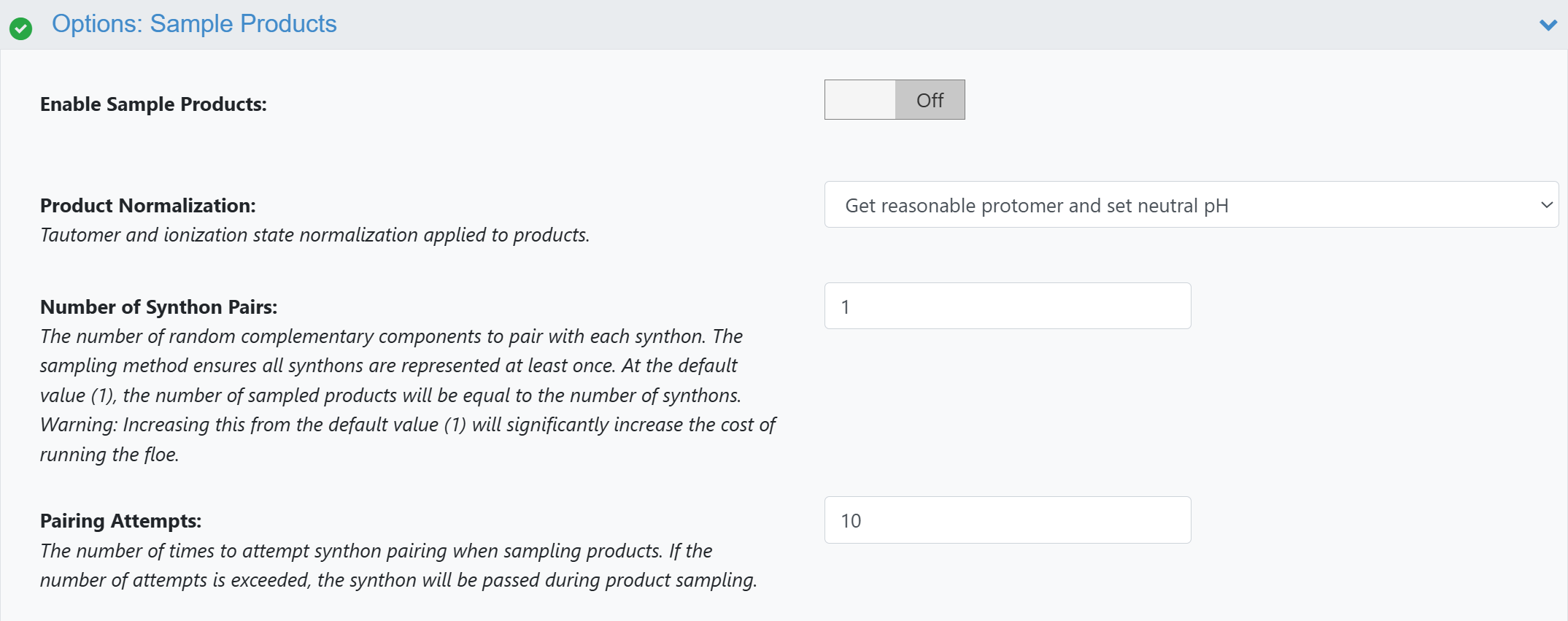
Figure 1. Turning off product sampling in ROCS X - Prepare 3D Library.
Note
You cannot turn product sampling off in the Multi-Stage ROCS X Preparation Floe.
The floe should run more quickly since it skips generating conformers for a product sample. Because the ROCS X 3D library no longer has an initial product sample to seed the model with during initialization, you need to generate a hit list for the query on your own. There are many tools for doing this on Orion (such as FastROCS Plus or Molecule Search). See the FastROCS Plus with Freeform Analysis tutorial and the documentation for the Molecule Search page.
As an example, this guide uses Molecule Search to generate a hit list for the query in Tutorial Query Mol: TNKS2–4l33–pdb-ligand.
Find your query in the Data page and add it to active data by clicking the ‘+’ icon next to the resource name (the ‘+’ icon will become a green check, and the query should appear in Active Datasets in the Active Data Bar).
In the 3D & Analyze page, right-click on the Molecule field in the spreadsheet and select “Send to Molecule Search.” The Molecule Search page will open and load your query.
Run a 3D Similarity search using any of the prepared 3D Databases. For this example search, a database with <500,000,000 molecules should suffice. Also, set the Max Hits parameter to 10000. Click the “Search” button.
Depending on the size of the database, Molecule Search will take about a minute to complete. After the search, take some time to examine the results. When you are done, click the “Save as dataset” button to create a hit list for the search. This is the external hit list that replaces the Initial Sample Hitlist for seeding the model during initialization.
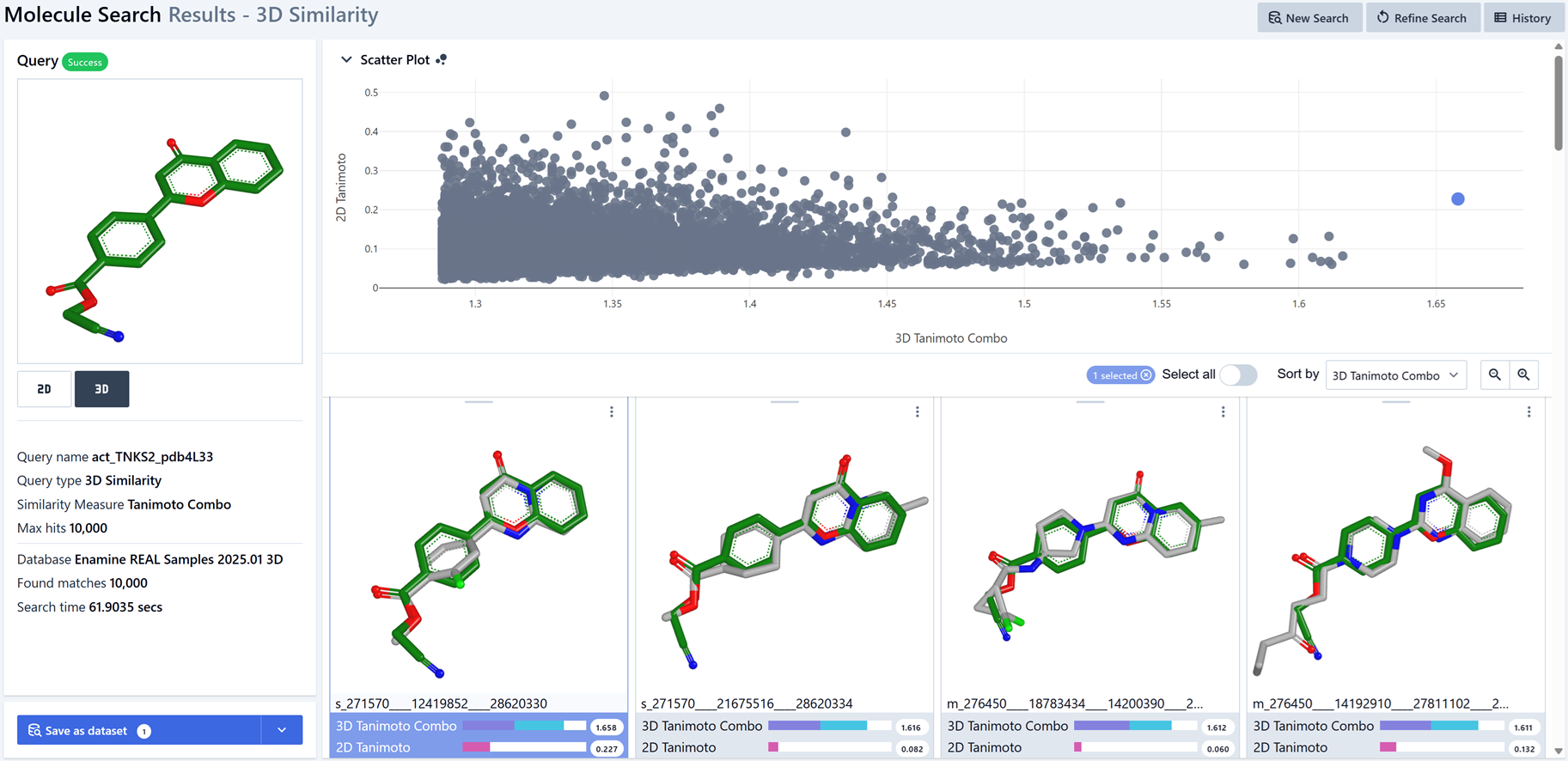
Figure 2. Running 3D Similarity Molecule Search on a query.
Now, when initializing a model for 3D search with the ROCS X - Initialize 3D Search Floe, use the parameters as in Figure 3 below. There are similar options to input an external hit list in the Multi-Stage ROCS X Search Floe.
Initialize with Sampled Products: Turn Off.
External Hit List: Use the Molecule Search hit list you saved above.
External Hit List Score Field: Make sure this matches the field on External Hit List for your scoring function. For Molecule Search - 3D Similarity searches, the default field name is “3D Tanimoto Combo”.
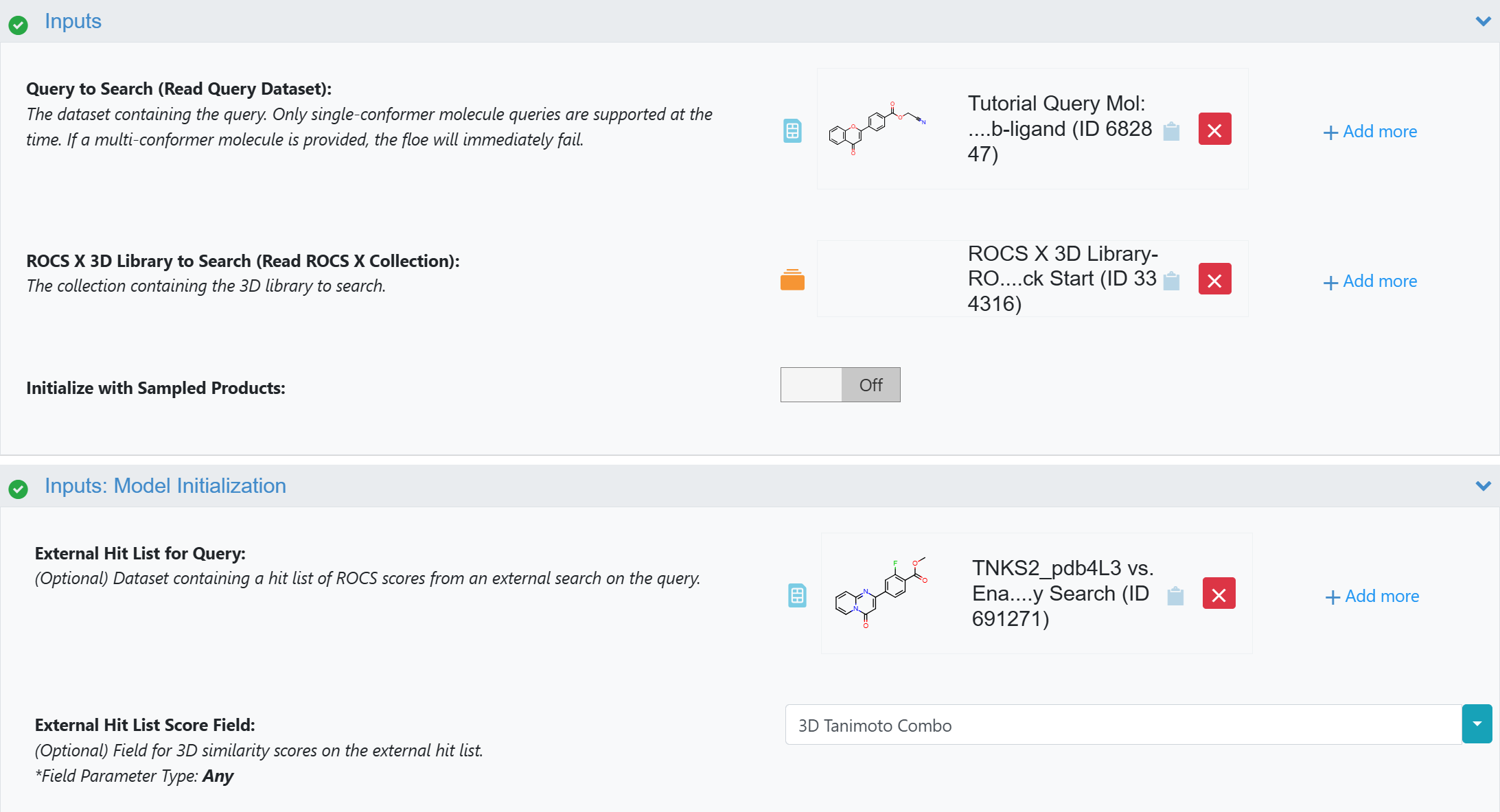
Figure 3. Seeding the model with an external hit list in the ROCS X - Initialize 3D Search Floe.
Note
If you’re initializing a model with a ROCS X 3D Library that doesn’t have a product sample, it doesn’t matter if you turn Initialize with Sampled Products Off. For best practices, since you’re not using the product sample to initialize the model, turn it Off.
When you run the floe, the initial FastROCS search on the query with the product sample will be skipped, and the external hit list will be used to seed the model. In Figure 4 of the introduction, this is equivalent to replacing the Initial Sample Search and Initial Sample Hit List with a Molecule Search and Molecule Search Hit List, respectively.