OEInterAdaptor
Attention
This API is currently available in C++ and Python.
class OEInterAdaptor : public OEMolAdaptor
The OEInterAdaptor class provides the ability to optimize a molecule or conformation with the use of internal coordinates (bond lengths, bond angle bonds and dihedral angles) defined by the internal Z-matrix.
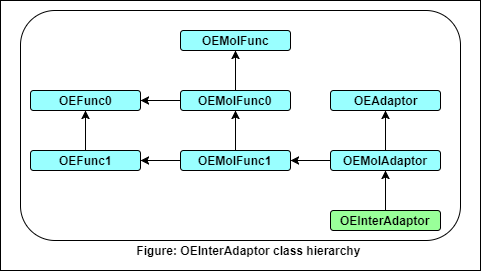
- The following methods are publicly inherited from OEAdaptor:
- The following methods are publicly inherited from OEFunc0:
- The following methods are publicly inherited from OEFunc1:
- The following methods are publicly inherited from OEMolFunc:
- The following methods are publicly inherited from OEMolAdaptor:
- The OEInterAdaptor class defines the following public methods:
Constructors
OEInterAdaptor(OEMolFunc1 &, bool dihedrals=true, bool own=false)
The molecule function used for function evaluation must be provided as the first argument to the constructor. The second argument allows to fix all dihedral angles. By default all dihedral angles are optimized. The third argument specifies whether the OEInterAdaptor object takes ownership of the memory of the molecule function instance. By default the OEInterAdaptor instance does not take ownership of the molecule function, so the OEInterAdaptor destructor does not delete the molecule function instance. If ownership of the molecule function is transferred to the OEInterAdaptor instance, the molecule function’s delete operator will be called in the OEInterAdaptor destructor.
Default and copy constructors.
AdaptGrad
bool AdaptGrad(double *ga, const double *g) const
Transforms the set of Cartesian gradients passed as a second argument to the method, into a set of internal coordinates gradients and copies them into the array passed as a first argument.
Set
bool Set(const OESystem::OEUnaryPredicate<OEChem::OEAtomBase>&,
unsigned int functype = OEFuncType::InterAdapt)
bool Set(const OESystem::OEUnaryPredicate<OEChem::OEBondBase>&,
unsigned int functype = OEFuncType::InterAdapt)
Those two methods assign atom and bond predicates which are used to partially restrict molecules
during optimization. First method specifies atoms which are to be held fixed in their starting
positions. Second method is used to fix dihedral angles a1-a2-a3-a4 defined by a middle bond a2-a3.
Both methods Set must be called before the method
derived from Setup.