DockingReport
Overview
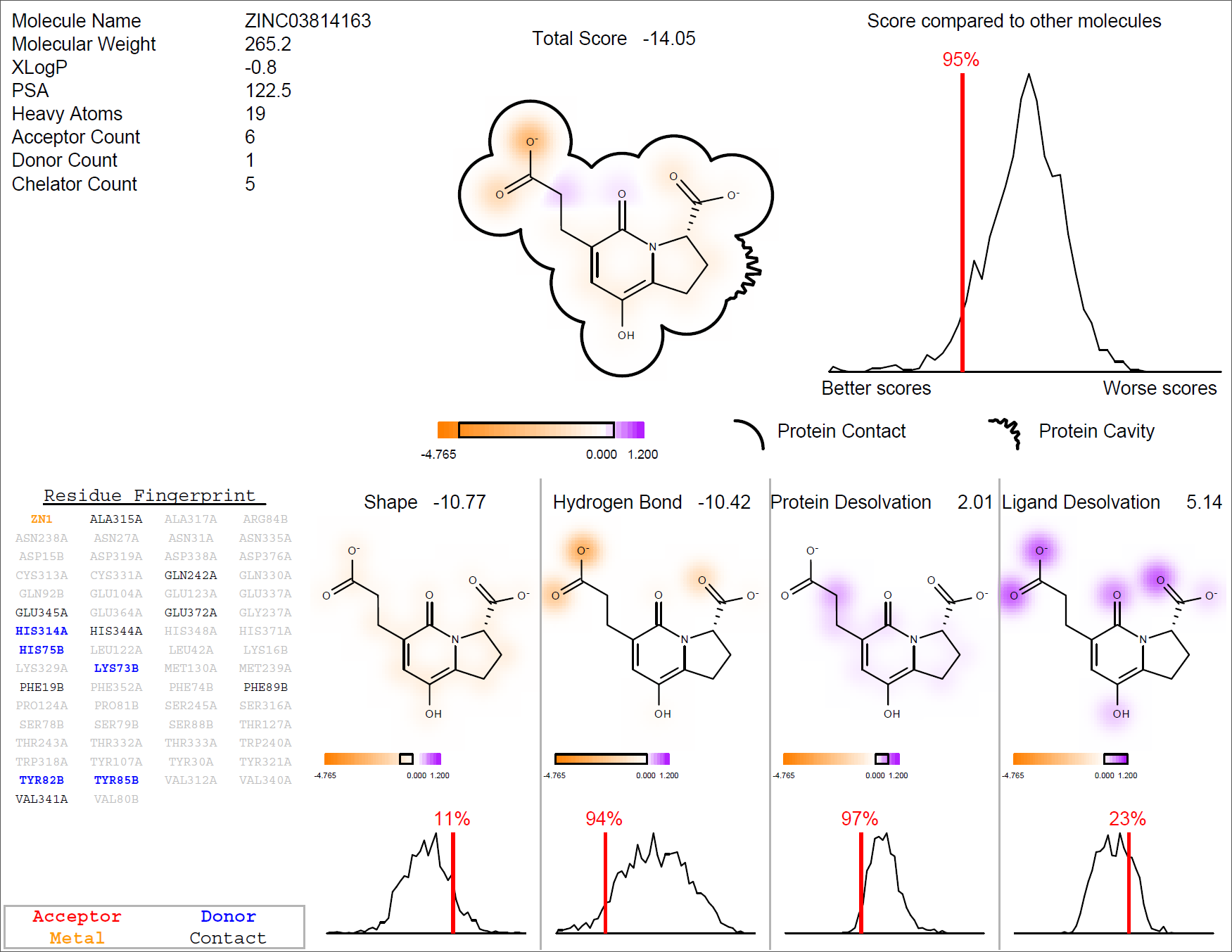
DockingReport is a utility program that creates a PDF report for one or more molecules docked by FRED or HYBRID. The PDF report lists includes a 2D depiction of each molecule, a breakdown of the score by atom, a breakdown of the score components by atom, a comparison of the molecule’s score compared to the other docked molecules, SD data tagged to the molecule and other general molecule properties (e.g., molecular weight). DockingReport also includes a ‘Residue Fingerprint’ which highlights which residues in the receptor site the ligand is interacting with (greyed out residues are residues in the site that other ligands interact with, but the current ligand does not).
|
Example dock report |
Example Commands
Creating a report for named molecules
Creates a report for 2 molecules blockbuster1 and blockbuster2 that were docked by FRED.
Input files
fred_docked.oeb.gz
Molecules docked by FRED including two molecules named blockbuster1 and blockbuster2 respectively.
receptor.oedu
The receptor file the molecules in docked.oeb.gz were docked to.
Command line
prompt> docking_report -docked_poses fred_docked.oeb.gz \ -receptor receptor.oedu \ -names blockbuster1 blockbuster2
Output files
docking_report.pdf
PDF dock report for blockbuster1 and blockbuster2.
Creating a report for molecules identified by SMILES
Creates a report for molecules, docked by HYBRID, that have the same canonical smiles as molecules in a specified file.
Input files
hybrid_docked.oeb.gz
Molecules docked by HYBRID.
receptor1.oedu and receptor2.oedu
The receptor files HYBRID used to dock the molecules in docked.oeb.gz.
interesting_ligands.sdf
Set molecules to create reports for. The molecules in this file will be converted into canonical SMILES, and a report will be generated for each molecule in docked.oeb.gz that has the same canonical SMILES as any one of the molecules in interesting_ligands.sdf
Command line
prompt> docking_report -docked_poses hybrid_docked.oeb.gz \ -receptor receptor1.oedu receptor2.oedu \ -smiles_file interesting_ligands.sdf
Output files
docking_report.pdf
PDF dock report for all molecules in docked.oeb.gz that have the same canonical SMILES as one of the molecules in interesting_ligands.sdf.
Command Line Help
A description of the command line interface can be obtained by executing DockingReport with the –help option.
> docking_report --help
will generate the following output:
Help functions:
docking_report --help simple : Get a list of simple parameters
docking_report --help all : Get a complete list of parameters
docking_report --help defaults : List the defaults for all parameters
docking_report --help <parameter> : Get detailed help on a parameter
docking_report --help html : Create an html help file for this program
docking_report --help versions : List the toolkits and versions used in the application
Required Parameters
Input
- -docked_poses <filename>
A file with poses that were docked by either FRED or HYBRID.
[ Aliases = -poses ]
- -receptor <filename> [<filename> ...]
Receptor file the poses passed to
-docked_poseswere docked too. If the poses were docked with hybrid HYBRID using multiple receptor structures then the same set of receptor structures should be passed to this flag.[ Aliases = -rec ]
Optional Parameters
Input Options
- -names <molecule name> [<molecule name> ...] [No Default]
This option identifies one or more molecules passed to the
-docked_posesoption by name. Molecules specified with this flag will have a report generated for them.Note
Either this flag or
-smiles_filemust be specified.[ Aliases = -title ]
- -smiles_file <filename> [No Default]
This option identified molecules passed to the
-docked_posesoption by canonical SMILES. Molecules identified with this flag will have a report generated for them.Note
Either this flag or
-namesmust be specified.[ Aliases = -smiles ]
- -sd_tags <Tag> [<Tag> ...] [No Default]
This flag specifies a specific set of SD tagged data to include in the dock report. If this flag is not specified all the SD data will be include in the dock report.
Note
The dock report has a limited amount of space for SD data and can only include 10 pieces of SD data for each docked molecule (additional SD data will not appear in this report).
[ Aliases = -sdtags ]
Output Options
- -report_file <filename> [Default: docking_report.pdf]
Name of the docked report file to generate.
The file must be a PDF file (i.e., have a .pdf extension).
[ Aliases = -report ]