POSIT Theory
POSIT is a pose-prediction tool primarily based on the assumption that similar ligands bind similarly. Pose prediction is the process of determining the structure of a ligand bound in the active site of a target protein. Pose prediction with POSIT assumes that the incoming ligand binds to the given protein and makes the best effort in correctly placing it in the active site. In conjunction to the predicted pose, POSIT also provides an estimate of confidence, in terms of an estimated probability, that the docked pose is within 2.0 Ångströms of the actual binding pose.
Note
Note that this probability is not the probability of binding, rather, the probability that the pose is correct given the ligand actually binds to the receptor.
POSIT consists of multiple Docking or Pose Prediction methods intentionally and automatically chooses the best method to use for any particular ligand based on the 2D (graph) and 3D (structure) similarity of the docked ligand to the bound ligand. The methods POSIT uses to dock are:
If there is no bound ligand in the receptor, the Fred method is used by default since it does not rely upon the pressence of the bound ligand.
Handling of isomerisms and chirality
Stereo, and notably nitrogen aniline stereo centers, are currently somewhat problematic for POSIT. Many crystal structures have flat geometries for some stereo centers due to time-averaging during data collection. This makes stereo centers appear to have flat geometries in the 3D coordinates.
Because the POSIT algorithm internally expands conformations during the flexible fitting procedure, the full molecule must be labeled with stereo - either in the 3D coordinate sense or the 2D coordinate sense. This means that some pdb ligand structures will unfortunately be failed by the POSIT algorithm.
To get around this, posit can be told to ignore nitrogen stereo during the conformer generation phase.
Note
Output structures of Posit are not guaranteed to have the same conformations of the input molecule. This is due to the fact that force-field minimization is occasionally performed during pose prediction.
Estimated Pose Probability
During a drug discovery campaign, thousands of small molecule inhibitors are made in the course of optimizing molecular properties. For projects that have X-Ray crystallographic (XRC) coordinates, structure-based designs help guide the medicinal chemistry efforts. In many cases XRC provides a detailed picture of the binding of a small-molecule inhibitor into the binding site.
Many techniques exist for pose-prediction and are well documented [Erickson-2004] . However, very few provide a probability that the generated pose is correct where correct is typically considered to be less than 2.0 Ångströms RMSD (root mean square distance) from experimental crystal structure. In fact, many docking scores such as Chemscore, Chemgauss3, PLP [Tuccinardi-2010] are not very correlated with correct ligand pose, and worse are not transferable between systems. The best docking score in one system may not even be close to the best docking score in another.
POSIT overcomes these issues by comparing predicted poses to observed bound ligands in related co-crystals. As the observed ligand becomes more similar to the the predicted pose, both the binding mode and the shape of the receptor pocket itself tends to become more similar.
The similarity measures being used are 2D path-based fingerprints and the 3D TanimotoCombo [Hawkins-2010] that compares shape and the Mills-Dean approximation of electrostatics [Mills-Dean-1996] . These similarity measures choose the most appropriate system to dock against (when multiple receptors are available) and provide a prediction of the quality of the result. The TanimotoCombo measure is agnostic of how the poses in question are generated; it can be used to validate and provide a pose prediction probability regardless of how the pose was generated.
POSIT probabilities were generated using a large test set containing many pose predictions and verified against an independent set of predictions that were then validated with X-Ray crystallography. It is important to note that POSIT does not give a probability of binding. Rather it gives a probability that if the ligand does actually bind, what is the likelihood of the POSIT pose being the actual pose.
Figure POSIT Probability MAP shows how the beliefs given by the 2D and 3D measures are combined into a probability of having a good pose. Remember that this probability has been generated from ligands that actually bind, hence, it is not a probability of binding.
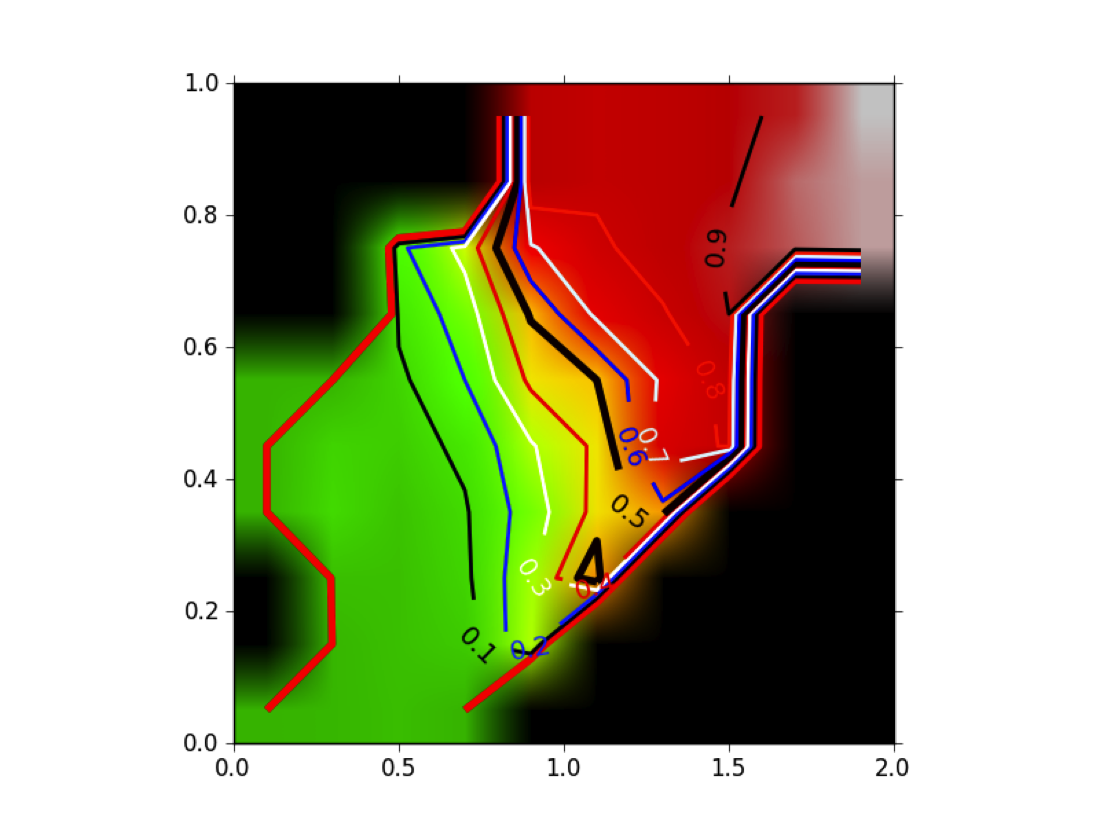
POSIT Probability Map: Given a 2D similarity (in this case the MACCS 166 descriptor set [Durant-2002]) and a 3D similarity (TanimotoCombo) posit computes a probability of finding the correct pose based on an analysis of historical and experimental data.
This result is different from the result shown in [Tuccinardi-2010] , where they reported that having a high TanimotoCombo to the known bound ligand did not dramatically increase the quality of the resulting pose (even for FRED). The reason is subtle: Tuccinardi et al were computing the highest TanimotoCombo that the two molecules could obtain, while Posit computes the actual, docked, in-place TanimotoCombo of the fitted pose. That is, if the docking algorithm produces an alignment of fit molecule to known bound ligand that overlaps with a given TanimotoCombo, one can look up the probability the docking was successful. In point of fact, Posit is specifically designed so that the docked pose obtains the highest TanimotoCombo score possible while simultaneously minimizing induced strain and maintaining interactions with the protein.
Categorically the data shown in POSIT Probability MAP, for each pose can be binned into the following results:
Result
Meaning
GREAT
Computed pose is likely (75%-100% probability) to be within 2.0 Å of experimentally-derived pose.
GOOD
Computed pose may be (50%-75% probability) to be within 2.0 Å of experimentally-derived pose.
MEDIOCRE
Take with a grain of salt (33%-50% probability)
POOR
Take with a huge grain of salt (<33% probability)
Additional MCS Constraints
Walter’s et al noted that a large portion of ligands bound to the same protein kinase share a large maximum common substructure (MCS). This was the basis for their CORES algorithm [Hare-2004] . Posit can optionally identify matching regions and use them as additional constraints during conformer generation. Posit performs the MCS match to the bound ligand by default.
On Clashes
The definition of clashes is somewhat problematic for purposes of pose prediction. In general, serious clashes where interpenetration with the protein should be avoided at all costs. However, when docking into a rigid protein that does not have the appropriate conformation, rigid docking ignores that fact that the active site may adopt a conformation suitable to the posed ligand.
Posit uses the definitions from OEClashInteractionHint to identify clashes. Posit allows the users to specify three allowable clash levels.
Allowed Clash
Description
noclashes (none)
No clashes are allowed.
hydrogen (mild clashes)
Only hydrogen clashes are allowed.
allclashes
All clashes are allowed.
Using Multiple Receptors
Using multiple receptors to dock or pose molecules has been shown to greatly increase the reliability of docking.
When multiple receptors are provided Posit chooses the appropriate reference system based on the 2D/3D similarity to the bound ligand. The best reference, in general, has the highest 2D/3D similarity of the input molecule to the chosen bound ligand. The similarity measure used here are similar to those used in determining the best docking method in Posit.
Receptor Flexibility
Incorporating multiple receptors is the simplest way to account for receptor flexibility during pose prediction. With multiple receptors, Posit would pick the best suitable conformer of the receptor for a particular docked ligand.
Pose generated from Posit with provided receptor conformers can still sometimes contain clashes. Option is available to trigger post pose prediction relaxation, that relaxes both the posed ligand and protein residues around it.
Note that optimization of a protein-ligand complex can sometimes be time consuming, especially if the target protein and associated components are large.
TanimotoCombo
Posit uses the TanimotoCombo measure to compare (and optimize) predicted and bound ligands. The TanimotoCombo measure is simply two separate Tanimoto measures added together. While most uses of Tanimoto have been to compare fingerprints together, there is a direct relation between the 1D fingerprint bit vector and 3D space:
The basic equation for a field Tanimoto between two fields A and B is:
In the case of Posit, the field in question can be thought of as field of voxel space. For \(Tanimoto_{shape}\), where A and B are now molecules: if two objects fill the same voxels, then the Tanimoto value is 1.0. If two objects overlap by half, the Tanimoto value is 0.5 and so on. (The term voxel is used for purposes edification, in actuality the volumes estimated using a fast approximate method)
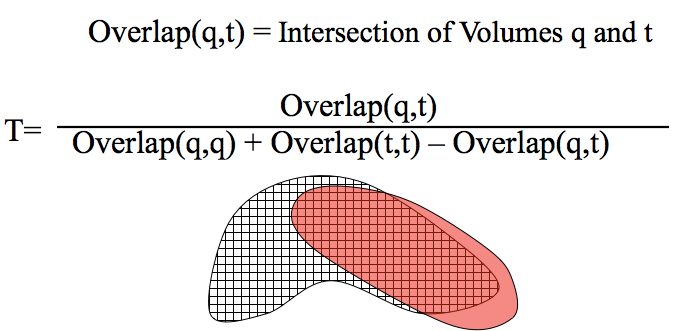
Voxel Representation of Shape: Similar to fingerprint bits in 1D, voxels can be used to represent 3D space and compared with the Tanimoto measure. The numerator Overlap(q,t) is essentially the volume of the intersection of q and t and the denominator Overlap(q,q) + Overlap(t,t) - Overlap(q,t) is essentially the volume of the union of q and t.
The field can also contain colored representations of chemistry. For example, if two voxels are colored as hydrogen bond donors and overlap, the \(Tanimoto_{color}\) increases.
Hence, TanimotoCombo is:
TanimotoCombo values range from 0 (no overlap) to 2.0 (full shape overlap and full color or chemistry overlap).