Protein and Ligand Preparation
In this section SPRUCE is used to prepare the proteins. You are free to use alternative methods to perform this and other “prep” tasks.
Note
If pre-prepared structures in .oedu fileformat from SPRUCE are available, these can
directly be used as input and the rest of this page can be skipped.
When using OEDesignUnits, note that they potentially include both solvent and
a bound ligand, depending on the structure. There are two flags that are crucial to keep
in mind, -excludeDUsolvent and -excludeDUligand, to control whether those two
components should be part of the calculation or not.
Good protein and ligand preparation is vital before running SZMAP or GamePlan. This consists of building partial sidechains, capping chains breaks, adding hydrogens, and assigning partial charges, and atomic radii.
The commands below are shown in a form appropriate for Linux and macOS. When you install SZMAP on Windows, a pre-configured version of the DOS command prompt is constructed and can be used to run any SZMAP program or utility without any extra setup. This window can be found under the Start menu in All Apps >> OpenEye-applications >> OpenEye-applications {version} Command line.
First, examine your structure in VIDA. VIDA can read gzipped structure files as-is and has the File >> Open Special >> From PDB menu command to fetch structures from the Protein Data Bank directly.
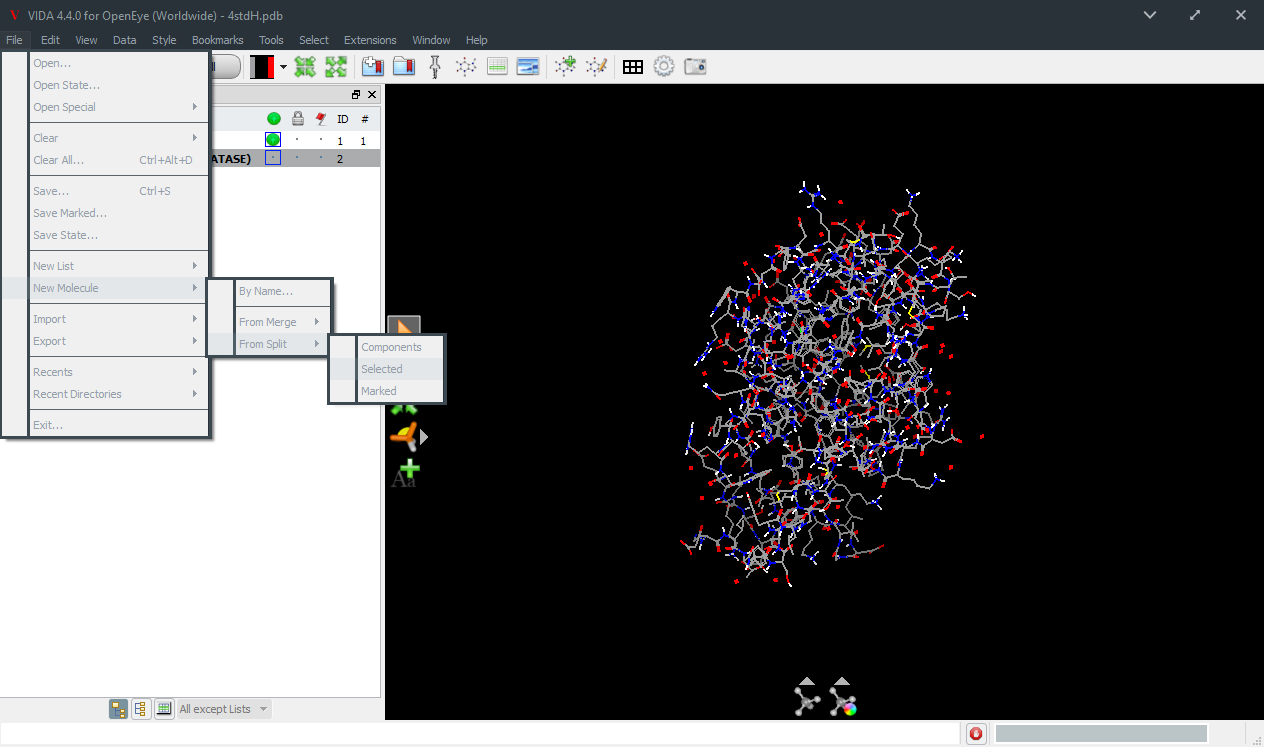
File >> New Molecule >> From Split >> Selected
The structure for 4STD is a trimer with the ligand some distance from the protein/protein interface. If you wish to delete detergents or other extraneous molecules, select everything you want to keep and make sure they are not selected. Then go to the menu item and split out the piece desired. File >> New Molecule >> From Split >> Selected. Alternatively, you may prefer to edit your molecule by hand as follows. If your structure file is gzipped PDB file, unzip it. Edit it to delete extra subunits, detergent, or other extraneous molecules using whichever text editor you prefer. It is not necessary to remove the waters at this stage, they wil be split out later. It is also not necessary to prune the connection table—any references to deleted atoms will be ignored
Since SZMAP and GamePlan require explicit hydrogen atoms on the molecules and most PDB structures do not include the hydrogen atoms, the next steps produce a protein structure with all the hydrogen atoms explicitly represented. There are many ways to do this. Here, we will use SPRUCE, the application will automatically generate biologically relevant units from the asymmetric unit (if needed), enumerates alternate locations, builds partial sidechains, caps chain breaks, places and optimizes protons including tautomer searching for ligands and co-factors. The alterante location enumeration and biological unit generation, means the application can output multiple structures based on the input unit.
> spruce -in 4std.pdb -prefix szmap_
szmap_4STD_A__DU__BFS_A-173.oedu
In this case the application wrote one structure into three files, the target/receptor structure, the automatically detected ligand, and a molecule containing the solvent. In these files the molecules have already had charges and radii assigned. If the ligand molecule is < 100 atoms, it will have AM1BCC charges assigned along with Zap9 radii, otherwise it will have MMFF charges assigned and Bondi radii. Protein molecules will have AmberFF94 charges assigned for any standard amino acid residue and AM1BCC for nonstandard residues. The function will fall back to assign the entire protein MMFF94 charges if there is a failure. Nucleic acids will have MMFF94 charges assigned. Water molecules are assigned SZMAP water parameters, described further in the The Szmap TK Standard Water Model chapter of the SZMAP toolkit.